 |
 |
- Search
| Genomics Inform > Volume 10(2); 2012 > Article |
Abstract
Dyslipidemia, mainly characterized by high triglyceride (TG) and low high-density lipoprotein cholesterol (HDL-C) levels, is an important etiological factor in the development of cardiovascular disease (CVD). Considering the relationship between childhood obesity and CVD risk, it would be worthwhile to evaluate whether previously identified lipid-related variants in adult subjects are associated with lipid variations in a childhood obesity study (n = 482). In an association analysis for 16 genome-wide association study (GWAS)-based candidate loci, we confirmed significant associations of a genetic predisposition to lipoprotein concentrations in a childhood obesity study. Having two loci (rs10503669 at LPL and rs16940212 at LIPC) that showed the strongest association with blood levels of TG and HDL-C, we calculated a genetic risk score (GRS), representing the sum of the risk alleles. It has been observed that increasing GRS is significantly associated with decreased HDL-C (effect size, -1.13 ± 0.07) compared to single nucleotide polymorphism combinations without two risk variants. In addition, a positive correlation was observed between allelic dosage score and risk allele (rs10503669 at LPL) on high TG levels (effect size, 10.89 ± 0.84). These two loci yielded consistent associations in our previous meta-analysis. Taken together, our findings demonstrate that the genetic architecture of circulating lipid levels (TG and HDL-C) overlap to a large extent in childhood as well as in adulthood. Post-GWAS functional characterization of these variants is further required to elucidate their pathophysiological roles and biological mechanisms.
Dyslipidemia is an important etiological factor in the development of cardiovascular disease (CVD), which is the leading cause of deaths for adults. Dyslipidemia is mainly characterized by elevated levels of triglycerides (TGs), increased low-density lipoprotein cholesterol (LDL-C), and reduced high-density lipoprotein cholesterol (HDL-C) [1].
In the past few years, genome-wide association studies (GWASs) and meta-analyses have identified common genetic variants that contribute to blood lipid phenotypes in Caucasian-based populations [2-8]. In addition, we have convincingly mapped 16 loci for major lipoprotein concentrations (TG, HDL-C, and LDL-C) in a two-stage association study in individuals of East Asian ancestry [9].
Given the growing global epidemic of metabolic syndrome, including obesity and type 2 diabetes (T2D), lipoprotein metabolism disorders have become an important health problem in adulthood but also have emerged as an increasingly prevalent risk factor in childhood [10]. Although many GWASs and meta-analyses have provided susceptibility genes or loci for lipid-related traits from multiple large population-based cohorts of adults, it is of great importance to evaluate genetic predisposition at an early age. However, little is known about their genetic effects in a younger population. Moreover, most identified variants have shown modest effects and have explained only a small proportion of heritable components for disease risk [11]. Nevertheless, when multiple genetic markers are considered together, literature-based genetic risk scores might be useful in improving the identification of the genetic contribution in those at high risk of disease [12-14]. Therefore, the aim of this study was to consolidate the associations of GWAS-based candidate variants with blood lipid concentrations in a childhood obesity study and to evaluate genetic risk scores (GRS) for lipid-related risk phenotypes.
Korean Association Resource (KARE) study was collected through the Korean Genome Epidemiologic Study project - in total, 10,038 people, 40-69 years old, living in Ansung and Ansan, from 2001-2002. For more information, the study has already been reported [9, 15]. The Health Examinee cohort (HEXA) shared project is a community-based epidemiological cohort of 1,200,000 subjects (aged, 40-69) who were recruited for baseline examination from 2004 to 2009. To share the controls, 4,302 subjects were randomly selected to participate between 2006 and 2007. The BioBank Japan study was approved by the ethical committees in the Institute of Medical Science, the University of Tokyo, and the Center for Genomic Medicine, RIKEN. Subjects for the Health2 cohorts for the replication study were selected from another community-based cohort sample provided by the Health2 study, as described previously [15]. We examined 7,861 selected from the 8,500 participants (aged, 40-69). The students aged 9 and 12 years were recruited from Gwacheon City and Gyeonggi Province between April and June 2010. This study was conducted as part of the Korean Children-Adolescents Study (KoCAS), which has been monitored yearly since their entry into elementary school at age 7 in Gwacheon City or fourth grade at age 10 in Seoul and Gyeonggi Province, Korea. Subjects who were enrolled in a specific diet program or were taking any medications known to affect appetite were excluded from the study. The study protocol was approved by the institutional review board of Seoul-Paik Hospital, Inje University, and the Korea Center for Disease Control and Prevention. Informed consent was obtained from the children's parents.
KARE samples were genotyped using Affymetrix Genome-Wide Human SNP array 5.0 (Affymetrix, Santa Clara, CA, USA) and processed by Bayesian Robust Linear Modeling using the Mahalanobis Distance (BRLMM) genotyping algorithm [16]. The HEXA shared sample was genotyped using Affymetrix Genome-Wide Human SNP array 6.0 and processed with the Birdseed Genotyping Algorithm [17]. The BioBank Japan study was genotyped using the Illumina Human610-Quad BeadChip (Illumina Inc., San Diego, CA, USA). A childhood obesity study was genotyped using the Illumina Omni1-Quad BeadChip. Individuals were excluded as follows: genotyping call rate, sex inconsistency, heterozygosity, identity-by-state value, and any kinds of tumor. For more information, the study has been reported [15].
For the replication study in Health2 (7,861), we performed a genotype assay using the TaqMan reaction for four single nucleotide polymorphisms (SNPs) (rs2074356, rs16940212, rs12708980, rs599839) and the GoldenGate assay (Illumina Inc.) for nine SNPs (rs780092, rs10503669, rs2001945, rs603446, rs12686004, rs11216126, rs12229654, rs519113, rs12654264, rs2738446). To analyze the quality control, we conducted duplicate genotyping using 1-2.5% of samples. For further association analyses, concordance rates were satisfied with duplicates of over 99% and a genotype success rate of over 98%.
In each GWAS data point, imputation analysis was performed using the IMPUTE reference HapMap Asian (Japanese [JPT]/Han Chinese [CHB]) population panel, based on NCBI build 36 and dbSNP build 129. Of these, in each cohort, we dropped SNPs with a posterior probability score < 0.90, high genotype information content (info < 0.5), Hardy-Weinberg equilibrium (HWE) (p < 1 × 10-7), SNP missing rate > 0.1, and minor allele frequency (MAF) < 0.01. In the former analysis, only imputed SNPs that satisfied the genomewide SNP quality control criteria were considered.
In performing the association tests, SNPs were analyzed with PLINK (http://pngu.mgh.harvard.edu/~purcell/plink/) and SAS version 9.1 (SAS institute Inc., Cary, NC, USA). Individuals who were on lipid-lowering therapy and hypertensive medications were excluded for dyslipidemia-related phenotypes, such as TG, HDL-C, and LDL-C. LDL-C concentrations were calculated with Friedewald's formula [18]. Missing values were assigned for individuals with TG over 400 mg/dL. The dyslipidemia-related phenotypes were tested by multivariate linear regression analysis in an additive genetic model (1-d.f.), including age, sex, and recruitment area as covariates. TG and HDL-C were transformed prior to analysis using natural log transformation to remove skewing. LDL-C was normally distributed, and no transformations were required. A GRS was examined using multivariate linear regression analysis, adjusting for age, sex, and recruitment area.
The clinical characteristics and statistics of the variables for each study sample are described in Table 1. Our previous large-scale GWAS identified 9 novel and 24 known genetic loci influencing metabolic traits in East Asians [9]. In an attempt to determine whether well-established lipid-related variants contribute to significant and reproducible genetic factors for susceptibility to blood lipid concentrations in a childhood obesity study, we first examined the associations between individual genetic variants and lipid-related phenotypes using linear regression analysis.
Of these 16 SNPs for blood lipid traits, rs10503669 in LPL (genotyped SNP in a childhood obesity study) was significantly associated with TG and HDL-C, respectively. In addition, rs16940212 in LIPC (linkage disequilibrium-based proxy SNP) was associated with HDL-C (Table 2). The direction of these associations was consistent with our previous GWAS [9].
Having two loci (rs10503669 at LPL and rs16940212 at LIPC) showing the strongest association with TG and HDL-C, we next investigated whether the cumulative allelic dosage of risk alleles contributes to the quantitative variation of lipoprotein concentrations in the KARE study using a method of calculating a GRS. We evaluated the joint effects of the best-associated SNPs at the two loci showing evidence of association with HDL-C (rs10503669 at LPL and rs16940212 at LIPC). We calculated a GRS, representing the sum of the risk alleles, and observed that increasing GRS was significantly associated with decreased HDL-C (effect size, -1.13 ± 0.07) compared to SNP combinations without two risk variants (Fig. 1). In addition, a positive correlation was observed between allelic dosage score and risk allele (rs10503669 at LPL) for high TG levels (effect size, 10.89 ± 0.84) (Fig. 2).
Blood lipid levels are an important etiological contributor to CVD. CVD is the leading cause of death among individuals with T2D, in which lipid abnormalities are characterized by hypertriglyceridemia, reduced levels of HDL-C, and elevated levels of LDL-C. These lipid-related traits, including ~14-54% for TG, ~34-42% for HDL-C, and ~28-50% for LDL-C, are highly heritable in familial T2D or diabetic patients [19, 20]. These estimates indicate that genetic variants play important roles in explaining inter-individual variations in blood lipid levels.
Although systematic reviews have reported relationships between childhood obesity and adult CVD risk [21, 22], there is not a shred of evidence that blood lipid status in adults is an independent or dependent risk factor in a childhood obesity group. Early childhood obesity might be the best time for dyslipidemia intervention, including screening, early detection, and management. The aim of this study was to consolidate the associations of 16 GWAS-based candidate variants with blood lipid concentrations across the lifespan.
In GWAS-based candidate approaches, the well-known LPL gene was significantly associated with low TG and high HDL-C levels in a childhood obesity study, respectively. This finding is well matched to the negative correlation between blood TG and HDL-C concentrations [23]. In addition, the LIPC gene was further observed in decreased HDL-C. In analyses of the joint effects of these variants, we found that GRSs on the two risk alleles (rs10503669 at LPL and rs16940212 at LIPC) were associated with a cumulative effect of TG and HDL-C levels. However, these facts indicate that currently identified risk variants might have low discriminatory ability and modest genetic contribution to disease prediction [24].
Lipoprotein lipase (LPL) plays an important role in lipid metabolism by hydrolyzing TGs of circulating chylomicrons and very low-density lipoproteins. In addition, LPL is expressed in the brain regions that are functionally relevant to learning, memory, and other cognitive functions. Many GWASs and meta-analyses have shown that variations in the well-known LPL gene for multiple lipid-related traits are implicated in multiethnic populations [2, 3, 9, 25-28]. An increasing number of studies have suggested an association of LPL gene variants with the risk of cerebrovascular as well as CVD [29]. Recent genome-wide investigations identified schizophrenia-associated loci in the chromosome 8p22 region, including LPL [30-32]. It was suggested that LPL is an attractive candidate gene that might be involved in the potential role of lipid metabolism in schizophrenia. Stages of early childhood development can be taken into account when considering a diagnosis of mental illness [33].
Hepatic lipase (LIPC) gene encodes hepatic triglyceride lipase, which is expressed in liver. LIPC has a dual function as a triglyceride hydrolase and ligand/bridging factor for receptor-mediated lipoprotein uptake. A GWAS has identified a possible role of LIPC in advanced age-related macular degeneration (AMD) [34]. AMD is an eye condition affecting the central part of the retina. Although a condition that is commonly associated with the elderly, macular degeneration affects younger people through a rare inherited condition [35]. Based on a lack of direct relationships between HDL-C and AMD risk, the LIPC association may not be the result of an effect on HDL-C levels, but it could represent a pleiotropic effect as a functional component, such as a lipoprotein transporter, underlying the biological mechanisms involving the cholesterol pathway.
Taken together, our findings demonstrate that the genetic architecture of circulating lipid levels (TG and HDL-C) overlap to a large extent in childhood as well as in adulthood. A genetic risk assessment by incorporating two risk alleles suggests encouraging evidence that identified genetic risk variants for lipid phenotypes have cumulative effects on lipid concentrations. Post-GWAS functional characterization of these variants is further required to elucidate the pathophysiological role and biological mechanisms.
Acknowledgments
This work was supported by grants from the Korea Centers for Disease Control and Prevention (4845-301, 4851-302, 4851-307) and an intramural grant from the Korea National Institute of Health (2011-N73005-00).
References
1. Meagher EA. Addressing cardiovascular disease in women: focus on dyslipidemia. J Am Board Fam Pract 2004;17:424–437. PMID: 15575034.


2. Kristiansson K, Perola M, Tikkanen E, Kettunen J, Surakka I, Havulinna AS, et al. Genome-wide screen for metabolic syndrome susceptibility Loci reveals strong lipid gene contribution but no evidence for common genetic basis for clustering of metabolic syndrome traits. Circ Cardiovasc Genet 2012;5:242–249. PMID: 22399527.



3. Tan A, Sun J, Xia N, Qin X, Hu Y, Zhang S, et al. A genome-wide association and gene-environment interaction study for serum triglycerides levels in a healthy Chinese male population. Hum Mol Genet 2012;21:1658–1664. PMID: 22171074.


4. Aulchenko YS, Ripatti S, Lindqvist I, Boomsma D, Heid IM, Pramstaller PP, et al. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nat Genet 2009;41:47–55. PMID: 19060911.



5. Kathiresan S, Melander O, Guiducci C, Surti A, Burtt NP, Rieder MJ, et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nat Genet 2008;40:189–197. PMID: 18193044.



6. Kathiresan S, Willer CJ, Peloso GM, Demissie S, Musunuru K, Schadt EE, et al. Common variants at 30 loci contribute to polygenic dyslipidemia. Nat Genet 2009;41:56–65. PMID: 19060906.



7. Willer CJ, Sanna S, Jackson AU, Scuteri A, Bonnycastle LL, Clarke R, et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat Genet 2008;40:161–169. PMID: 18193043.



8. Diabetes Genetics Initiative of Broad Institute of Harvard and MIT, Lund University, and Novartis Institutes of BioMedical Research. Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI, et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 2007;316:1331–1336. PMID: 17463246.


9. Kim YJ, Go MJ, Hu C, Hong CB, Kim YK, Lee JY, et al. Large-scale genome-wide association studies in East Asians identify new genetic loci influencing metabolic traits. Nat Genet 2011;43:990–995. PMID: 21909109.


10. Bradfield JP, Taal HR, Timpson NJ, Scherag A, Lecoeur C, Warrington NM, et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat Genet 2012;44:526–531. PMID: 22484627.



11. Zuk O, Hechter E, Sunyaev SR, Lander ES. The mystery of missing heritability: genetic interactions create phantom heritability. Proc Natl Acad Sci U S A 2012;109:1193–1198. PMID: 22223662.



12. Liu S, Song Y. Building genetic scores to predict risk of complex diseases in humans: is it possible? Diabetes 2010;59:2729–2731. PMID: 20980472.



13. Qi L, Ma J, Qi Q, Hartiala J, Allayee H, Campos H. Genetic risk score and risk of myocardial infarction in Hispanics. Circulation 2011;123:374–380. PMID: 21242481.



14. Vaarhorst AA, Lu Y, Heijmans BT, Dollé ME, Böhringer S, Putter H, et al. Literature-based genetic risk scores for coronary heart disease: the Cardiovascular Registry Maastricht (CAREMA) prospective cohort study. Circ Cardiovasc Genet 2012;5:202–209. PMID: 22373668.


15. Cho YS, Go MJ, Kim YJ, Heo JY, Oh JH, Ban HJ, et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat Genet 2009;41:527–534. PMID: 19396169.


16. Rabbee N, Speed TP. A genotype calling algorithm for affymetrix SNP arrays. Bioinformatics 2006;22:7–12. PMID: 16267090.


17. Yeung JM, Sham PC, Chan AS, Cherny SS. OpenADAM: an open source genome-wide association data management system for Affymetrix SNP arrays. BMC Genomics 2008;9:636. PMID: 19117518.



18. Johnson R, McNutt P, MacMahon S, Robson R. Use of the Friedewald formula to estimate LDL-cholesterol in patients with chronic renal failure on dialysis. Clin Chem 1997;43:2183–2184. PMID: 9365406.


19. Malhotra A, Wolford JK. American Diabetes Association GENNID Study Group. Analysis of quantitative lipid traits in the genetics of NIDDM (GENNID) study. Diabetes 2005;54:3007–3014. PMID: 16186406.


20. Elbein SC, Hasstedt SJ. Quantitative trait linkage analysis of lipid-related traits in familial type 2 diabetes: evidence for linkage of triglyceride levels to chromosome 19q. Diabetes 2002;51:528–535. PMID: 11812765.


21. Lloyd LJ, Langley-Evans SC, McMullen S. Childhood obesity and adult cardiovascular disease risk: a systematic review. Int J Obes (Lond) 2010;34:18–28. PMID: 19434067.


22. Beilin L, Huang RC. Childhood obesity, hypertension, the metabolic syndrome and adult cardiovascular disease. Clin Exp Pharmacol Physiol 2008;35:409–411. PMID: 18307730.


23. Gaw A. HDL-C and triglyceride levels: relationship to coronary heart disease and treatment with statins. Cardiovasc Drugs Ther 2003;17:53–62. PMID: 12843687.


24. Cornelis MC, Qi L, Zhang C, Kraft P, Manson J, Cai T, et al. Joint effects of common genetic variants on the risk for type 2 diabetes in U.S. men and women of European ancestry. Ann Intern Med 2009;150:541–550. PMID: 19380854.



25. Franceschini N, Carty C, Bůzkova P, Reiner AP, Garrett T, Lin Y, et al. Association of genetic variants and incident coronary heart disease in multiethnic cohorts: the PAGE study. Circ Cardiovasc Genet 2011;4:661–672. PMID: 22042884.



26. Sarzynski MA, Jacobson P, Rankinen T, Carlsson B, Sjöström L, Carlsson LM, et al. Association of GWAS-based candidate genes with HDL-cholesterol levels before and after bariatric surgery in the Swedish obese subjects study. J Clin Endocrinol Metab 2011;96:E953–E957. PMID: 21430028.


27. Lettre G, Palmer CD, Young T, Ejebe KG, Allayee H, Benjamin EJ, et al. Genome-wide association study of coronary heart disease and its risk factors in 8,090 African Americans: the NHLBI CARe Project. PLoS Genet 2011;7:e1001300. PMID: 21347282.



28. Waterworth DM, Ricketts SL, Song K, Chen L, Zhao JH, Ripatti S, et al. Genetic variants influencing circulating lipid levels and risk of coronary artery disease. Arterioscler Thromb Vasc Biol 2010;30:2264–2276. PMID: 20864672.



29. Munshi A, Babu MS, Kaul S, Rajeshwar K, Balakrishna N, Jyothy A. Association of LPL gene variant and LDL, HDL, VLDL cholesterol and triglyceride levels with ischemic stroke and its subtypes. J Neurol Sci 2012;318:51–54. PMID: 22541364.


30. Need AC, Ge D, Weale ME, Maia J, Feng S, Heinzen EL, et al. A genome-wide investigation of SNPs and CNVs in schizophrenia. PLoS Genet 2009;5:e1000373. PMID: 19197363.



31. Ma X, Deng W, Liu X, Li M, Chen Z, He Z, et al. A genome-wide association study for quantitative traits in schizophrenia in China. Genes Brain Behav 2011;10:734–739. PMID: 21679298.


32. Lewis CM, Levinson DF, Wise LH, DeLisi LE, Straub RE, Hovatta I, et al. Genome scan meta-analysis of schizophrenia and bipolar disorder, part II: Schizophrenia. Am J Hum Genet 2003;73:34–48. PMID: 12802786.



33. Xie C, Wang ZC, Liu XF, Wang L, Yang MS. Association between schizophrenia and single nucleotide polymorphisms in lipoprotein lipase gene in a Han Chinese population. Psychiatr Genet 2011;21:307–314. PMID: 21862952.


34. Neale BM, Fagerness J, Reynolds R, Sobrin L, Parker M, Raychaudhuri S, et al. Genome-wide association study of advanced age-related macular degeneration identifies a role of the hepatic lipase gene (LIPC). Proc Natl Acad Sci U S A 2010;107:7395–7400. PMID: 20385826.



35. Harkewicz R, Du H, Tong Z, Alkuraya H, Bedell M, Sun W, et al. Essential role of ELOVL4 in very long chain fatty acid synthesis and retinal function. J Biol Chem 2011 12 24 [Epub]. http://dx.doi.org/10.1074/jbc.M111.256073.



Fig. 1
Combined effects of risk alleles on high-density lipoprotein cholesterol (HDL-C). Allelic dosage scores are plotted on the X axis. The histogram (Y axis on left, grey bars) represents the percentage of individuals in each risk allele category. The histogram (Y axis on right, black bars) represents mean HDL-C concentrations. (A) Eight single nucleotide polymorphisms (SNPs) including the risk alleles (rs10503669 at LPL and rs16940212 at LIPC). (B) Six SNPs without the risk alleles (rs10503669 at LPL and rs16940212 at LIPC).
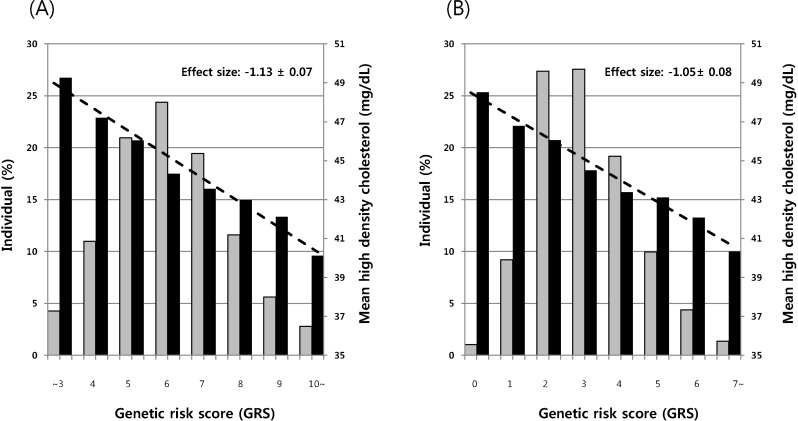
Fig. 2
Combined effects of risk alleles on triglyceride (TG). Allelic dosage scores are plotted on the X axis. The histogram (Y axis on left, grey bars) represents the percentage of individuals in each risk allele category. The histogram (Y axis on right, black bars) represents mean TG concentrations. (A) Five single nucleotide polymorphisms (SNPs) including the risk allele (rs10503669 at LPL). (B) Four SNPs without the risk allele (rs10503669 at LPL).

Table 1.
Descriptive statistics of variables for each study cohort
| Study |
Overall meta |
Replication |
|||
|---|---|---|---|---|---|
| KARE | HEXA-shared control | BBJ | Health2 | Childhood obesity | |
| Sample size | 8,842 | 3,703 | 15,764 | 7,861 | subjects |
| Age (y) | 52.22 ± 8.92 | 53.21 ± 8.33 | 63.7 ± 10.5 | 56.58 ± 7.85 | 10.83 ± 1.43 |
| Male/Female | 4,183/4,689 | 1,651/2,052 | 10,400/5,364 | 3,124/4,647 | 232/252 |
| TG (mg/dL) | 162.44 ± 104.53 | 123.29 ± 91.4 | 151.2 ± 108.6 | 157.26 ± 108.94 | 76.43 ± 43.69 |
| HDL-C (mg/dL) | 44.66 ± 10.1 | 54.61 ± 13.27 | 51.8 ± 14.6 | 46.34 ± 11.24 | 58.93 ± 12.34 |
| LDL-C (mg/dL)a | 115.65 ± 32.14 | 119.04 ± 31.39 | 126 ± 34 | 127.70 ± 34.57 | 95.80 ± 24.02 |
Table 2.
Association between SNPs and lipid-related traits in the population with discovery sets and replication study
| Trait | SNP ID | CHR | Gene | Coded allele (Freq) |
Overall meta |
Replication |
||||
|---|---|---|---|---|---|---|---|---|---|---|
| Effect size (beta ± SEM) | p-value | Meta-analysis heterogeneity p-value | Effect size (beta ± SEM) | p-value | Proxy | |||||
| TG | rs780092 | 2 | GCKR | G (0.33) | -0.05 ± 0.005 | 4.58E-27 | 7.55E-01 | -0.018 ± 0.033 | 5.78E-01 | |
| rs2286276 | 7 | TBL2-MLXIPL | T (0.10) | -0.065 ± 0.008 | 1.44E-15 | 3.51E-02 | -0.062 ± 0.048 | 1.95E-01 | ||
| rs10503669 | 8 | LPL | T (0.12) | -0.086 ± 0.007 | 6.84E-39 | 8.77E-01 | -0.121 ± 0.044 | 6.82E-03 | ||
| rs2001945 | 8 | C (0.42) | 0.041 ± 0.004 | 1.43E-20 | 9.42E-01 | 0.028 ± 0.031 | 3.72E-01 | rs2980853a | ||
| rs603446 | 11 | ZNF259-APOA1/C3/A4/A5-BUD13 | T (0.23) | -0.088 ± 0.004 | 1.81E-86 | 5.90E-01 | 0.002 ± 0.033 | 9.62E-01 | rs1145208a | |
| HDL-C | rs10503669 | 8 | LPL | T (0.12) | 0.0423 ± 0.003 | 8.04E-43 | 3.13E-02 | 0.046 ± 0.018 | 8.48E-03 | |
| rs12686004 | 9 | ABCA1 | T (0.21) | -0.022 ± 0.003 | 2.18E-18 | 2.93E-01 | -0.01 ± 0.015 | 4.95E-01 | ||
| rs11216126 | 11 | C (0.20) | 0.032 ± 0.003 | 2.60E-34 | 9.52E-01 | -0.01 ± 0.014 | 4.45E-01 | rs11021969b | ||
| rs12229654 | 12 | CUX2 | G (0.14) | -0.028 ± 0.003 | 3.41E-23 | 9.02E-01 | 0.017 ± 0.012 | 1.63E-01 | rs991817b | |
| rs2074356 | 12 | C12orf51 | T (0.15) | -0.035 ± 0.003 | 6.95E-37 | 6.90E-01 | 0.008 ± 0.015 | 5.95E-01 | rs12423268b | |
| rs16940212 | 15 | LIPC | T (0.34) | 0.022 ± 0.002 | 1.21E-24 | 5.91E-02 | 0.034 ± 0.013 | 7.74E-03 | rs16940223a | |
| rs12708980 | 16 | CETP | C (0.10) | -0.04 ± 0.004 | 1.81E-28 | 8.55E-01 | -0.028 ± 0.021 | 1.89E-01 | ||
| rs519113 | 19 | PVRL2-TOMM40-APOE | C (0.15) | -0.018 ± 0.003 | 8.04E-11 | 7.03E-02 | -0.024 ± 0.018 | 1.81E-01 | ||
| LDL-C | rs599839 | 1 | CELSR2-PSRC1-SORT1 | G (0.06) | -5.225 ± 0.579 | 1.84E-19 | 1.15E-04 | 2.06 ± 4.113 | 6.17E-01 | |
| rs12654264 | 5 | HMGCR | T (0.48) | -2.711 ± 0.291 | 1.21E-20 | 4.89E-02 | 0.128 ± 1.798 | 9.43E-01 | rs3846662a | |
| rs651007 | 9 | ABO | A (0.26) | 2.203 ± 0.384 | 9.78E-09 | 9.88E-02 | 1.048 ± 1.98 | 5.97E-01 | rs579459a | |
| rs2738446 | 19 | LDLR | C (0.14) | 2.951 ± 0.420 | 2.02E-12 | 4.38E-01 | 3.374 ± 2.515 | 1.80E-01 | rs688a | |
- TOOLS
-
METRICS

-
- 7 Crossref
- 14,009 View
- 56 Download





