Possibility of the Use of Public Microarray Database for Identifying Significant Genes Associated with Oral Squamous Cell Carcinoma
Article information
Abstract
There are lots of studies attempting to identify the expression changes in oral squamous cell carcinoma. Most studies include insufficient samples to apply statistical methods for detecting significant gene sets. This study combined two small microarray datasets from a public database and identified significant genes associated with the progress of oral squamous cell carcinoma. There were different expression scales between the two datasets, even though these datasets were generated under the same platforms - Affymetrix U133A gene chips. We discretized gene expressions of the two datasets by adjusting the differences between the datasets for detecting the more reliable information. From the combination of the two datasets, we detected 51 significant genes that were upregulated in oral squamous cell carcinoma. Most of them were published in previous studies as cancer-related genes. From these selected genes, significant genetic pathways associated with expression changes were identified. By combining several datasets from the public database, sufficient samples can be obtained for detecting reliable information. Most of the selected genes were known as cancer-related genes, including oral squamous cell carcinoma. Several unknown genes can be biologically evaluated in further studies.
Introduction
Despite recent advances in surgical, radiation, and chemotherapeutic treatment protocols, the prognosis of oral squamous cell carcinoma (OSCC) remains mournful, with an approximate 50% 5-year mortality rate from disease or associated complications [1]. Therefore, the identification of biological markers is essential to make progress in detecting malignancy at an early stage and developing novel therapies [2].
Microarray datasets that are created for the same research purposes in different laboratories have accumulated rapidly. The results from different datasets are often inconsistent due to the utilization of different platforms, sample preparations, or various technical variations. If we could combine such datasets by adjusting for systematic biases that exist among different datasets derived from different experimental conditions, the power of statistical tests would be improved by the increase in sample size [3].
In OSCC, although lots of microarray-based studies have been conducted to provide insights into gene expression changes, most of these studies have contained insufficient samples for detecting reliable information using statistical analysis [4, 5]. Therefore, this study attempted to combine several datasets in the public database for detecting significant genes.
We used two small microarray datasets of OSCC for this study, which were based on the same platform but had different expression scales. These two datasets were combined after discretization, because a previous study showed that classification could be improved using combined datasets after discretization [3]. After combining datasets, we used chi-square test for identifying the significant genes. Chi-square test has been used commonly to detect differentially expressed genes after discretization of expression intensities in the microarray experiment.
In this study, gene expression ratios of two datasets were transformed with their ranks for each dataset. Next, the transformed datasets were combined, and a nonparametric statistical method was applied to the combined dataset to detect informative genes. Finally, we showed that most of the selected genes were known to be involved in various cancers, including OSCC.
Methods
Dataset
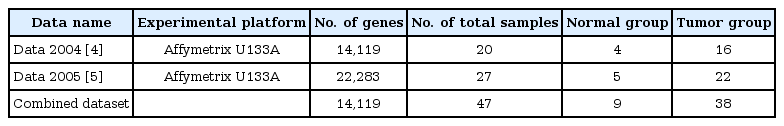
Two microarray datasets were used for this study. We acquired these datasets from a public database (Gene Expression Omnibus, GEO). One was the expression dataset of 16 tumors and 4 normal tissues from 16 patients, using Affymetrix U133A gene chips (Affymetrix, Santa Clara, CA, USA). The other microarray dataset consisted of expression profiles of 22 tumors and 5 normal tissues. These two datasets were experimented on under the same platform, Affymetrix U133A. The datasets are summarized in Table 1.
Process for combining datasets
For combining datasets, gene expression ratios are rearranged in order of expression ratios by each gene in each dataset, and the ranks are matched with the corresponding experimental group. If the experimental groups are homogenous, the ranks within the same experimental group would be neighboring. The process of discretization of gene expressions is summarized in the following steps [3]:
Rank the gene expression ratios within a gene for each dataset.
List in order of the ranks, and assign the order of gene expressions to the corresponding experimental groups.
Summarize the result of (2) in the form of a contingency table for each gene.
Combine the contingency tables that have been summarized for each dataset.
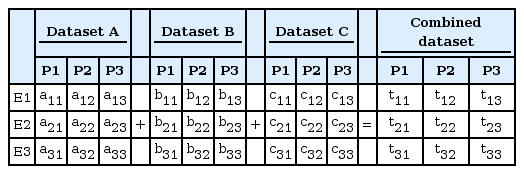
When there are three datasets to be combined, the datasets can be added as a single entry, as shown in Table 2, after the transformation of each dataset by rank.
Identification of significant genes from a combined dataset
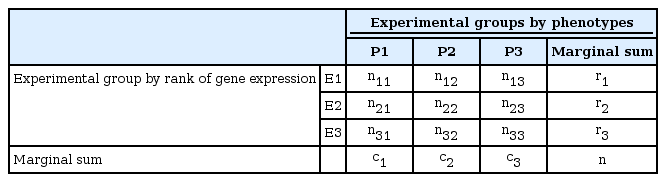
After the summarization of gene expression ratios in the form of a contingency table for each gene, as shown in Table 3, a nonparametric statistical method was applied to the datasets for independence testing between gene expression patterns and experimental groups. The test statistics are calculated as follows for each gene:
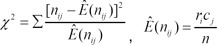
When the sample size is small - generally Ê(nij) less than 5 - Fisher's exact test is recommended rather than chi-square test.
The significant genes can be selected by an independence test between the phenotypes and gene expressions using this type of summarized dataset. ci and ri represent the marginal sums of the ith column and row, respectively. nij is the number of experiments belonging to Ei and Pj, and n represents the total number of experiments.
Results
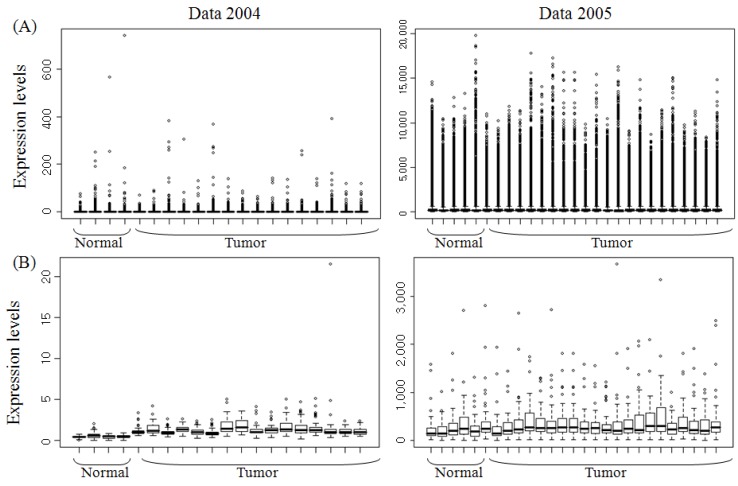
The clinical information and expression levels of two datasets are summarized in Table 4 and Fig. 1. Subgroup and sex were similarly distributed in the two datasets. The distributions of other factors were not included.
The scale of expression levels in the two datasets was different; the expression values of Data 2004 ranged from 0.01 to 740, and those of Data 2005 were from 0.1 to 19,773. The expression patterns of the two datasets can be explored in Fig. 1.
Lots of outliers are shown in Fig. 1A in the two datasets containing whole gene sets. However, in subsets of significant genes, the expression ranges got narrow, and the outliers were decreased (Fig. 1B). The expressions of tumor tissues in Data 2004 were upregulated and varied compared with normal tissues. If there was no outlier with a maximum value in the 14th tumor tissue in Data 2004, the expressions of the two different groups would be clearly distinguished. Any clear differences in expression were not shown between the two groups in Data 2005.
Upregulated 51 genes in oral squamous cell carcinoma
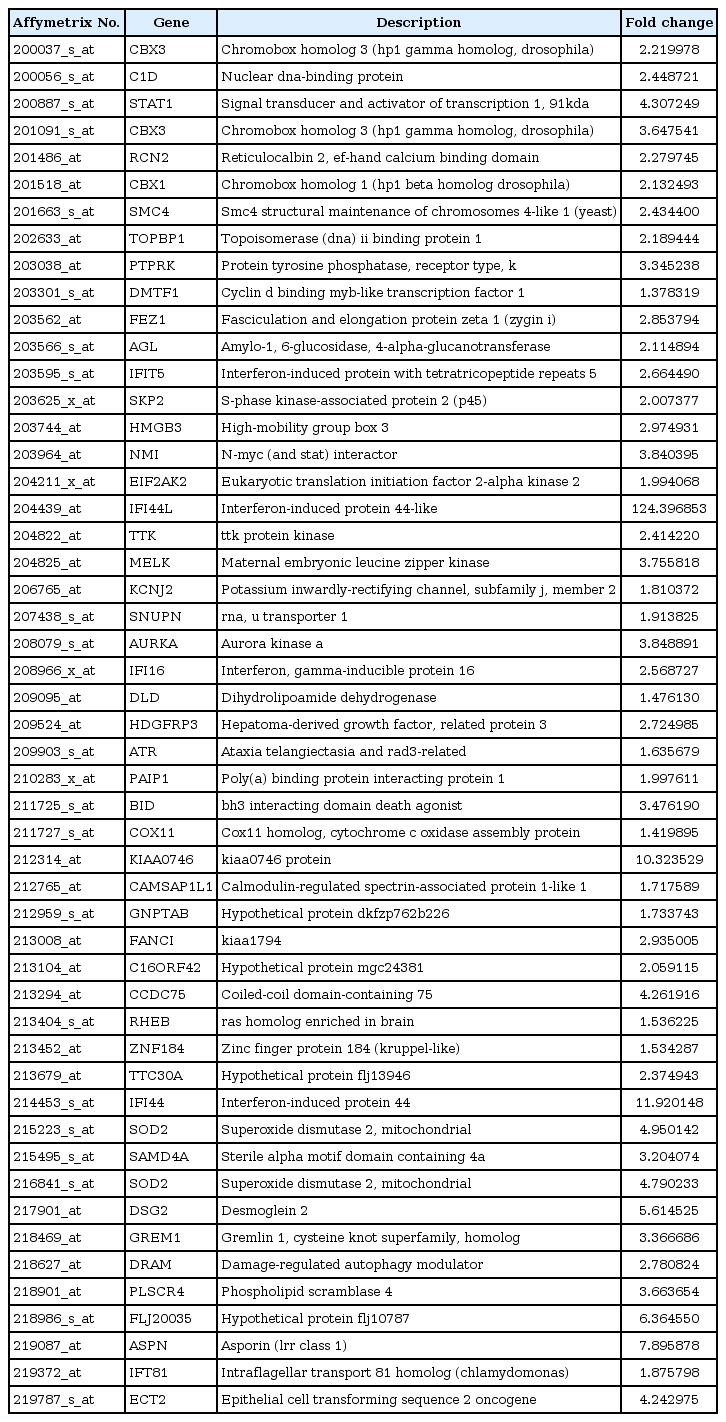
To identify differently expressed genes between normal and tumor tissues, we performed chi-square test using a combined microarray dataset. Fifty-one significant genes were selected from a combined dataset with p-value less than 0.005, which were upregulated in OSCC tissues. The significance level can be controlled, and more genes can be selected with a lower significance level. These selected genes are summarized in Table 5.
Many genes among the selected genes were known as cancer-related genes. STAT1 [6], SKP2 [7], IFI16 [8], RHEB [9], FIF44 [10], SOD2 [11, 12], and GREM1 [11] are related to OSCC. Table 6 [13-56] summarizes the previous studies that have published the relations of selected genes with cancer.
Expression pattern of the identified genes
To investigate whether the different experimental groups could be classified with significant genes, an unsupervised hierarchical clustering method was applied to the significant gene set (Fig. 2).
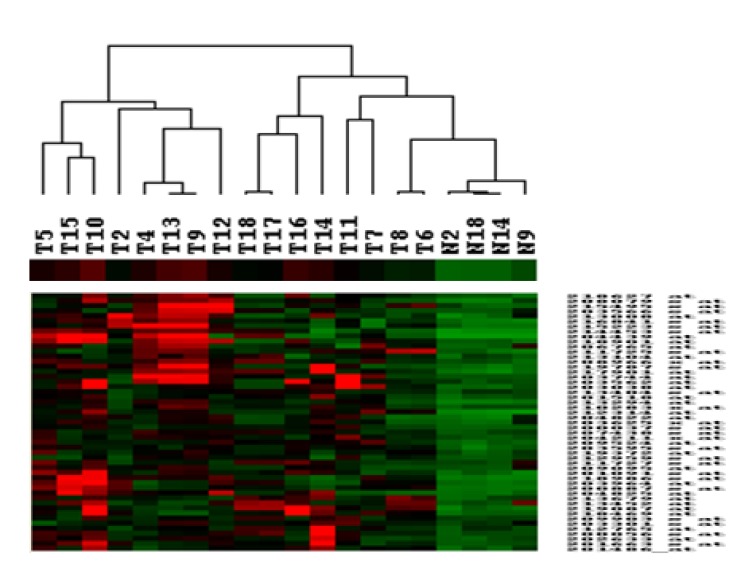
Expression patterns of the selected 51 genes. These genes were upregulated in oral squamous cell carcinoma tissues, and normal and tumor groups were clearly classified with these genes.
The normal group consisted of 4 tissues and showed significantly lower expression levels when compared with the tumor group. In Fig. 2, we investigated the classification availability of the identified genes in Data 2004, not in a combined dataset, because the two datasets have different expression scales.
Network analysis
Based on all identified genes, new and expanded pathway maps and connections and specific gene-gene interactions were inferred, functionally analyzed, and used to build on the existing pathway using the Ingenuity Pathway Analysis (IPA) knowledge base [57].
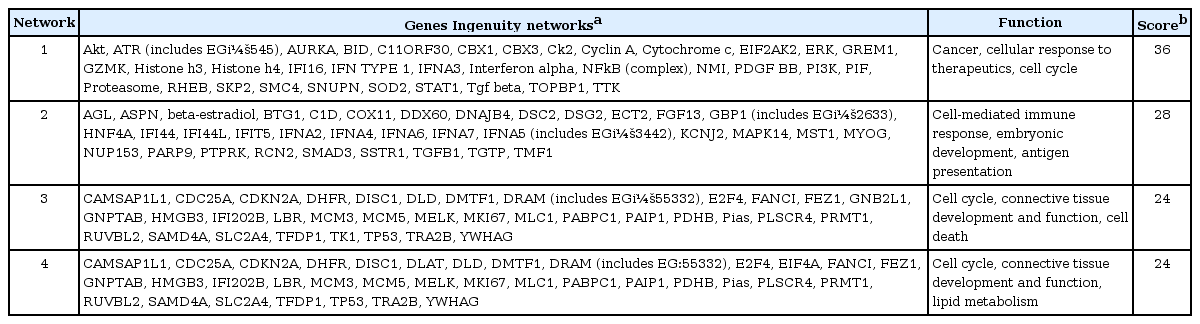
To generate networks in this work, the knowledge base was queried for interactions between the identified genes and all other genes stored in the database. Four networks were found to be significant in OSCC. The network with the highest score (Network 1, score = 36) was generated, with 17 identified genes (Table 7, Fig. 3).
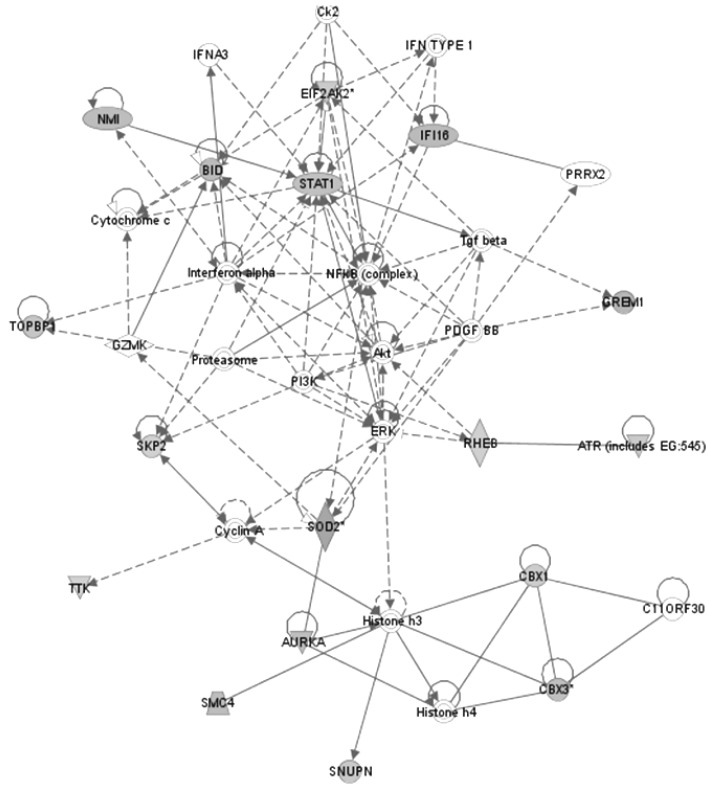
Network with the highest score (Network 1). Functional relationships between genes based on known interactions in Ingenuity Pathway Analysis (IPA) knowledge are described.
In the network diagram, STAT1 and SOD2 neighbored with NMI and AURKA, respectively. The expression levels of STAT1 and SOD2 could be expected to be related with those of NMI and SOD2. Actually, the expressions of STAT1 and SOD2 were strongly positively correlated with NMI (r = 0.95) and AURKA (r = 0.87), respectively.
Discussion
OSCC is associated with substantial mortality and morbidity [58]. To identify potential biomarkers for early detection of invasive OSCC, microarray experiments have been conducted, and these kinds of microarray datasets have accumulated rapidly in the public database. However, there are many datasets that include insufficient sample sizes for detecting significant genes by statistical analysis. Therefore, this study attempted to combine several microarray datasets from a public database to identify significant candidates as biomarkers.
In a microarray data analysis, the information from different datasets obtained under different experimental conditions may be inconsistent even though they are performed with the same research objectives. Moreover, even when the datasets are generated by the same platform, the data agreement may be affected by technical variations between laboratories. In such cases, it could be necessary to use a combined dataset after adjusting for the differences between such datasets for detecting the more reliable information. Combining datasets is especially useful in OSCC microarray datasets, because there are many datasets with insufficient sample sizes for analysis [4, 5, 59, 60].
For identifying significant genes classifying tumor and normal groups, we achieved two microarray datasets from a public database, GEO. They included 20 and 27 samples, and each sample size was unbalanced between the different groups. By combining these two datasets, the sample size was increased, and we had a sufficient sample size for statistical analysis, even though it was still unbalanced. When these datasets were combined, we used the rank of gene expression, because the scale of gene expression was different. In this study, we identified 51 significant genes from a combined dataset, and this number could be increased or decreased by the significance level (we used 0.005). The selected 51 genes were upregulated in tumor tissues. Many of the selected genes were proven to be cancer-related genes by previous studies.
SOD2 is associated with lymph node metastasis in OSCC and may provide predictive values for the diagnosis of metastasis [10]. Metastasis is a critical event in OSCC progression. An SOD2 variant has also been associated with increased breast cancer and ovarian cancer risk in previous studies [47, 61]. TopBP1 included eight BRCT domains (originally identified in BRCA1), and it was proposed as a breast cancer susceptibility gene [18, 62].
By semiquantitative reverse transcription PCR analysis, RHEB was shown to be upregulated in OSCC [9]. In salivary cancer, survival probability rates dropped when Skp2 was overexpressed [7]. Overexpression of Skp2 is associated with the reduction of p27 (KIP1) expression and may have a role in the progression of OSCC [25].
The expression of RCN2 was linearly related to the tumor mass increase, and its expression was increased in breast cancer [16]. PTPRK was proven as a candidate gene of colorectal cancer [19], and it is a functional tumor suppressor in Hodgkin lymphoma cells [20]. DMTF1 was shown to be amplified in adenocarcinoma of the gastroesophageal junction, residing at 7q21 by aCGH experiments [21]. FEZ1 was involved in ovarian carcinogenesis, and its reduction or loss could be an aid to the clinical management of patients affected by ovarian carcinoma [22]. It is also a known tumor suppressor gene in breast cancer and gastric cancer [23, 63].
Other ovarian cancer-related genes were NMI [27, 28] and FANCI [44]; breast cancer-related genes were COX11 [42], MELK [33], and FANCI [44] among the selected genes. MELK was known to be associated with shorter survival in glioblastoma [34].
TTK was associated with progression and metastasis of advanced cervical cancers after radiotherapy [29, 30]. It might also be a relevant candidate as a new target in cancer therapy, since it plays relevant roles in mitotic progression and the spindle checkpoint [31, 32]. Aurora kinase A (AURKA) was associated with skin tumors [36] and colorectal cancer [37, 38].
In previous studies, OSCC-related genes among the selected genes were STAT1 [14], SKP2 [7, 25], IFI16 [8], RHEB [9], IFI44 [64], SOD2 [10-12], and GREM1 [11]. The gene set, which has not been proven as OSCC-related genes until now, could be expected to be possibly proven as OSCC-related genes by biological evaluation.
In this study, we identified significant genes related with OSCC from two microarray datasets in a public database. For this, we transformed microarray datasets using ranks of gene expressions with different expression scales, even though they were constructed under the same experimental conditions. This method could be useful when using multiple datasets that are created for the same research purpose, By combining these accumulated datasets, we can detect more reliable information due to the increased sample size. It saves time and money and avoids repeating experiments.
Acknowledgments
This work was supported by the Priority Research Centers Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Education, Science and Technology (2011-0031396).