 |
 |
- Search
| Genomics Inform > Volume 20(2); 2022 > Article |
|
Abstract
The rise of newer coronavirus disease 2019 (COVID-19) variants has brought a challenge to ending the spread of COVID-19. The variants have a different fatality, morbidity, and transmission rates and affect vaccine efficacy differently. Therefore, the impact of each new variant on the spread of COVID-19 is of interest to governments and scientists. Here, we proposed mathematical SEIQRDVP and SEIQRDV3P models to predict the impact of the Omicron variant on the spread of the COVID-19 situation in South Korea. SEIQEDVP considers one vaccine level at a time while SEIQRDV3P considers three vaccination levels (only one dose received, full doses received, and full doses + booster shots received) simultaneously. The Omicron variantŌĆÖs effect was contemplated as a weighted sum of the delta and Omicron variantsŌĆÖ transmission rate and tuned using a hyperparameter k. Our modelsŌĆÖ performances were compared with common models like SEIR, SEIQR, and SEIQRDVUP using the root mean square error (RMSE). SEIQRDV3P performed better than the SEIQRDVP model. Without consideration of the variant effect, we donŌĆÖt see a rapid rise in COVID-19 cases and high RMSE values. But, with consideration of the Omicron variant, we predicted a continuous rapid rise in COVID-19 cases until maybe herd immunity is developed in the population. Also, the RMSE value for the SEIQRDV3P model decreased by 27.4%. Therefore, modeling the impact of any new risen variant is crucial in determining the trajectory of the spread of COVID-19 and determining policies to be implemented.
Coronavirus disease widely known as coronavirus disease 2019 (COVID-19) is a new disease caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus that emerged from Wuhan, China [1] and spread rapidly across the world becoming a global pandemic in March 2020 [2]. The pandemic caught many governments and people by surprise which led to the implementation of unprecedented intervention policies like school and workplace closures, suspension of public transportation, international travel restrictions, and so forth [3], with aims of mitigation and suppression. The reason was to not overwhelm the unprepared healthcare systems and lower the number of cases until a pharmacological solution was found [4,5]. In addition, many pharmaceutical companies in partnership with government bodies launched one of the fastest vaccine development projects of our decade leading to the development of multiple vaccines [6,7]. The Coalition for Epidemic Preparedness Innovations (CEPI) worked with global health authorities, governments, and vaccine developers to support the development of vaccines against COVID-19 [8].
This pandemic also attracted the interest of researchers from different fields since it was a new disease and its transmission pathways and fatality were not yet known. From the outbreak to 4 February, 2022, researchers have contributed a lot in forecasting, and understanding the transmission dynamics, the fatality of SARS-CoV-2, and the evolution of the pandemic, to help in the fight against this new global problem. Among the researchers, statisticians, epidemiologists, and mathematicians contributed to formulating models to capture the transmission dynamics of COVID-19 and forecasting the evolution of the pandemic among different populations amidst government interventions. These mainly included statistical models [9-18], deep-learning models [19-24] and mathematical models [1,14,25,26].
Statistical models offer more precise models and deep-learning techniques are the key to high-quality predictive models [27]. However, both statistical and deep-learning models require real data to make predictions. But with mathematical models, a set of mathematical equations that mimic the current situation is written, and solving them for certain parameters provide information about the disease characteristics [28]. Some of their advantages include mathematical models representing the real situation of the problem being solved and they do not require all data to be available for it to be fitted as deductions from known information about the situation can be used. Also, they can handle sudden changes and complexity with ease. Since the start of the COVID-19 pandemic, mathematical models have been at the forefront of determining and forecasting the spread of COVID-19 and shaping government policies around the world [28].
A seminal paper in 1927 introduced the Susceptible, Infectious, and Recovered (SIR), a mathematical model for infectious diseases [29]. Since then, with advances in information technology and fast computing methods, many variations of the SIR model have been developed. Because mathematical models can easily be understood and definite conclusions about the COVID-19 outbreak can be made from them, Susceptible, Exposed, Infectious and Recovered (SEIR), a modification of SIR and a cascade of other modifications have been constructed and developed for predicting COVID-19 since its declaration as a global pandemic [30-42].
Subsequently, on 8 December, 2020, 272 days after COVID-19 was declared a global pandemic, vaccination started in the United Kingdom [43]. Since then, as of 4 February, over 61.34% of the world population has received at least one dose of a COVID-19 vaccine, and 21.54 million doses of vaccines are administered daily around the world [44]. However, since then new variants of the SARS-CoV-2 virus have appeared. These variants have different transmissibility rates, fatality, and morbidity. Furthermore, existing vaccines have differing efficacy levels against these emerging variants [45,46]. Governments must make decisions and revise policies while considering these new developments like the impact of vaccinations and emerging variants on the spread of COVID-19. However, statistical and deep-learning models would require real data in substantial amounts to perform any forecasting or prediction. On the other hand, these new developments can easily be modeled with little or no data with mathematical models.
For the Korean COVID-19 situation, many models were employed to forecast the future COVID-19 situation in the country amidst government social distancing policies. One paper used the SIR model with time-dependent parameters and deep learning to forecast the spread of COVID-19 in South Korea [47]. Another analysis utilized the SIR model with breakpoint information that allows change in transmission rate at the breakpoints was established [48]. Other uses of the SIR model or its modification used for the Korean population are found elsewhere [49-53]. A modification of the SEIR model that considers transmission rates between age groups and vaccination was also formulated for the Korean population [54]. In this model, five additional groups; quarantined Q, unprotected U, vaccinated V, protected P, and deceased D were added to the standard SEIR model making it the SEIQRDVUP model.
Since the SARS-CoV-2 virus is an RNA virus and lacks the mismatch repair mechanism, the virus replication process is accompanied by a high mutation rate, hence the rise of variants [55]. Common mutant variants include B.1.1.7, B.1.351, B.1.1.28.1, B.1.617.2 (Delta), and B.1.1.529 (Omicron), which have all spread rapidly worldwide. The mutations make the virus more contagious (fast-spreading) and difficult to eliminate [56]. However, the SEIQRDVUP model and other previous methods cannot catch the sudden increase in daily cases caused by newer variants with higher transmission rates compared to a previously dominant variant.
To solve this limitation, we formulated a modification of the SEIQRDVUP model to consider a weighted sum of delta and Omicron variantsŌĆÖ transmission rates based on variantsŌĆÖ proportions together. In addition, three vaccination levels (only one dose received, full doses received, and full doses+booster shots received) were considered by adding three more compartments of vaccination (V1, V2, and V3) and the removal of the above-mentioned U compartment due to the use of a transmission rate that includes the effect of vaccine efficacy thereby eliminating the ineffectively vaccinated group, U. So, the Omicron variantŌĆÖs effect was contemplated as a weighted sum of the delta and Omicron variantsŌĆÖ transmission rate. In this case, the Omicron variantŌĆÖs transmission rate is assumed to be a multiple of the deltaŌĆÖs transmission rate, as explained in detail in the Methods section. This study aims to examine how the Omicron variant will affect the COVID-19 situation in Korea with our proposed SEIRQDVP and SEIRQDV3P models. The SEIQRDVP considers only one vaccination level at a time using only one vaccination compartment. However, the SEIQRDV3P considers all three vaccination levels simultaneously.
Mathematical methods can be used for the prediction and forecasting of COVID-19 transmission [57-60]. Here, we proposed the SEIQRDVP model, and its flowchart is shown in Figure 1 below. The susceptible group S is the group of unvaccinated and uninfected people that can still be infected by the infectious group. The vaccinated group V is a group of people vaccinated and can still be infected by the infectious group but with a lower transmission rate. In this case, the efficacy of the vaccine is multiplied by the transmission rate. If a host in S or V group gets infected, this host becomes a host of the exposed group, E. After the incubation period, a host of E can infect S or V groups, which means that a host of E becomes a host of I, the infectious group. When a host of I is determined to be infected, a host will be isolated and becomes a host of an isolated group, Q, and loses the ability to infect others. An isolated host will be recovered or be dead and moves to group R or D, which are the recovered group and deceased group. Group P is the insusceptible group that has immunity. The following differential equations represent the SEIQRDVP model:
where ╬▓ is the transmission rate, e is vaccine efficacy, f is the mortality rate, ╬▒,╬│,k and w are the duration periods from respective previous compartment to the next compartment, N is the total population, and ╬│ is the isolation duration.
Previously determined model parameters from literature, ╬║,╬▒,╬│,f,e, and used in our analysis are listed in Table 1. We assumed that the vaccinated host gets immunity 42 days after their first vaccination which means that 1/Žē is assumed to be 42 [57]. In Fig. 1, v is provided by daily vaccinated cases. Consequently, the remaining parameter ╬▓ is the only unknown parameter estimated by the least-squares method. This process is done using Runge-Kutta fourth-order method and the lsqcurvefit toolbox in MATLAB [61]. In addition, the daily cases are divided into segments with the breakpoints of these segments being determined from the changing levels of the stringency index due to changing government policies. The stringency index was obtained from the Oxford COVID-19 Government Response Tracker (OxCGRT) dataset from the Blavatnik School of Government and the University of Oxford [62,63]. ╬▓ was estimated for each segment independent of other segments, therefore our proposed model included stringency index as a covariate.
Moreover, the vaccination group can be divided into three: vaccinated (first vaccination), fully vaccinated (second vaccination), and boosted (third vaccination). We call the model that fits the three vaccination levels simultaneously, the SEIQRDV3P model, and its flowchart is shown in Fig. 2. In this model, v1,v2,v3 are provided by daily vaccinated, daily fully vaccinated, and daily boosted cases. Also, the efficacy of vaccination for each vaccination group is differently provided with e1, e2, e3 and values are 0.75, 0.80, 0.85 [69].
Lastly, the proportion of Omicron variants was reflected in the above model as a change in transmission rate, ╬▓. With a transmission rate of delta variant as ╬▓D and transmission rate of Omicron variant as ╬▓O, we assumed that ╬▓O is multiple of ╬▓D, which means ╬▓O=k╬▓D with hyperparameter k. In our cases, we tried 1, 3, 5, and 7 as a value of the hyperparameter k, to track the recent rapid increase of the Omicron variant. The proportion of the Omicron variant in the population is modeled by the parameter w. The values of w lie between 0 and 1. The time-series variation of this parameter is known for both train and test data, but its variation for the coming days is unknown. So, a logistic function was fitted to predict the future behavior of w. Using past data on the proportion of the Omicron variant, the logistic function of the proportion of the Omicron variant against time was fitted by the least square method. Combining these results altogether, the final transmission rate became ╬▓O w+╬▓D (1-w) which can be simplified as ╬▓D {1+(k-1)w}. Since k and w are constants, the only parameter estimated is ╬▓D, and was estimated by the same method as the above models.
Information of daily cases, deaths, and the three vaccination levels used in the analysis was obtained from the Our World in Data website [70]. The daily recovered data is obtained from a web-based dashboard tracker of COVID-19 hosted by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University [71]. The proportion of cases because of the Omicron variant was gotten from GISAID, an Initiative dedicated to the tracking of virus variants from the influenza viruses and coronavirus [72,73]. This data was divided into train and test data. The training period was chosen from 20 September 2021 to 28 January 2022 since from this date (2021 September 20), the proportion of cases of delta variant had exceeded 90% of the cases. The test data period for prediction was from 29 January 2022 to 4 February 2022.
Modifications of the basic SEIR model to the SEIQR model, to the SEIQRDVUP model, and then to our proposed SEIQRDVP and SEIQRDV3P models were done and the models' performances were compared. For each model, using train data, time-dependent ╬▓(t) using the different models were estimated by the least-squares method (LSE). SEIQR, SEIQRDVUP, and SEIQRDVP modelsŌĆÖ results showed similar fitting with our proposed SEIQRDV3P model. We observed that except for the basic SEIR model and our SEIQRDV3P model, the other models had similar daily cases fitted curves.
Using test data, the prediction error of each model using actual confirmed cases and predicted confirmed cases from models was determined using root mean square error (RMSE). RMSE values for the SEIR, SEIQR, SEIQRDVUP, SEIQRDVP, and SEIQRDV3P models were calculated as 11,235, 5,079, 5,116, 5,115, and 5,101, respectively, as shown in Table 2. A general decrease in RMSE values with an increase in model complexity is observed. However, the difference in RMSE between SEIQR to SEIQRDV3P models is way smaller than the difference between SEIQR and SEIR models.
From the above result, SEIQRDV3P and SEIQR models had the lowest but almost similar prediction errors, despite large differences in the model structure. This could be because of the recent Omicron situation in Korea. As transmissibility between the delta variant, which was originally dominant in Korea, and the Omicron variant which is now the dominant variant differ greatly. The SEIQRDV3P model which considers three vaccination levels simultaneously was updated to reflect the different effects of each variant due to their different transmissibility rate. Using train data, time-dependent ╬▓D(t) was estimated by the LSE method. In this case, hyperparameter k was chosen as 3, 5, 7, which means that the transmission rate of the Omicron is 3, 5, 7 times of transmission rate of the delta. For each selected k, the best fitted daily cases curves are shown in Fig. 3. The model with k=1 corresponds to the original SEIQRDV3P model. The x-axis (time) includes both the training and testing period.
Also, RMSE values for each case were calculated as 5,101, 4,583, 4,200, and 3,705 for each value of hyperparameter k (1, 3, 5, and 7). As we include the effect of the Omicron variant in the SEIQRDV3P model, we can observe the dramatic decrease in RMSE values. Also, RMSE values decreased as the hyperparameter k increased, as shown in Table 2. This result implies that in a short period, the Omicron variant shows way larger transmissibility than the delta variant. Seven daysŌĆÖ prediction after the test data period, which is 5 February 2022 to 11 February 2022, is shown in Fig. 4.
Since the onset of the global COVID-19 pandemic, mathematical models have been at the forefront of forecasting the future pandemic situation hence policymaking by government bodies. Mathematical models are highly flexible and the impact of different scenarios on the transmission of COVID-19 can be incorporated and predicted, even with the unavailability of data. The mathematical compartmental SEIR model and many of its modifications have been developed.
Governments must revise their testing protocols, social distancing policies, and healthcare protocols with the emergence of each new variant, hence the need of modeling the impact of each emerging variant on the spread of COVID-19. Here, we proposed a modification of the published SEIQRDVUP model, the SEIQRDVP model which considers one vaccination group at a time, and the SEIQRDV3P model which models the 3 vaccination levels simultaneously and the impact of the Omicron variant. SEIQRDVP and SEIQRDV3P modelsŌĆÖ performance were compared to SEIQRDVUP and other known compartmental mathematical models SEIR and SEIQR models. Firstly, without considering the Omicron variant rate, our SEIQRDV3P model doesnŌĆÖt show much difference from other models contrasted here. This result implies that the SEIQRDV3P model cannot predict a rapid increase in daily COVID-19 cases without a previous increasing daily case pattern.
However, using a hyperparameter and a weighted sum of transmission rates between two variants, we were able to predict the rapid increase caused by the Omicron variant. Omicron rate considering weighted sum lowers the prediction error of the SEIQRDV3P model from 5,101.342 to 3,705.078 which is 27.4% less than the SEIQRDVUP model. Since the Omicron variant has a way larger transmission rate than delta or other previously known variants, it seems that daily incidences will keep increasing until herd immunity for the Omicron variant is formed in the population.
However, from January 2022, daily deaths, as well as severity, seem to have decreased considerably. This pattern can imply the low risk and mortality associated with the Omicron variant compared to the delta variant, or the impact of vaccination on the population. Therefore, before implementing the ŌĆśLiving with COVID-19ŌĆÖ policy in Korea [74], the prediction of deceased and serious patient cases should be preceded. This work can be done by developing the mortality rate in the SEIQRDV3P model to also consider the Omicron variantŌĆÖs mortality with the weighted sum method.
Considering that each variant has its different transmissibility rates, fatality, impact on vaccine efficacy, and morbidity, this generates different model parameter values making it difficult to model all current variants in one model. Therefore, each variant would require its model. Currently, using different model parameters for each variant remained a limitation of this study which we try to solve in the future. Also, SARS-CoV-2 has been known to affect age groups differently. Furthermore, the impact of variant and vaccination policies across different age groups of the population will be considered in our future studies.
With the appearance of new COVID-19 variants appearing after a few months, the fight to end the spread of SARS-CoV-2 even with vaccination has been greatly challenged. These new variants have a different fatality, transmission rate, and efficacy from currently available vaccines. Therefore, their effect on daily cases, deaths, and implemented non-pharmacological policies is of interest to governments and scientists. With the proposed SEIQRDV3P model we found out the new Omicron variant will cause a rapid rise in COVID-19 cases in South Korea for some time until herd immunity is developed in the population.
Notes
AuthorsŌĆÖ Contribution
Conceptualization: TP. Data curation: JO. Formal analysis: JO. Funding acquisition: TP. Methodology: JO. Investigation: JO, CA. Visualization: JO, CA. Supervision: TP. Project administration: TP. Writing - original draft preparation: CA, JO. Writing - review and editing: TP, CA, JO.
Acknowledgments
This research was supported by the Bio and Medical Technology Development Program of the National Research Foundation (NRF) funded by the Korean government (MSIT) (No. 2021M3E5E3081425).
Fig.┬Ā2.
Flowchart of SEIQRDV3P model. Parameters for V1,V2,V3 to E are (1-e1)╬▓, (1-e2)╬▓,(1-e3)╬▓ like Fig. 1.
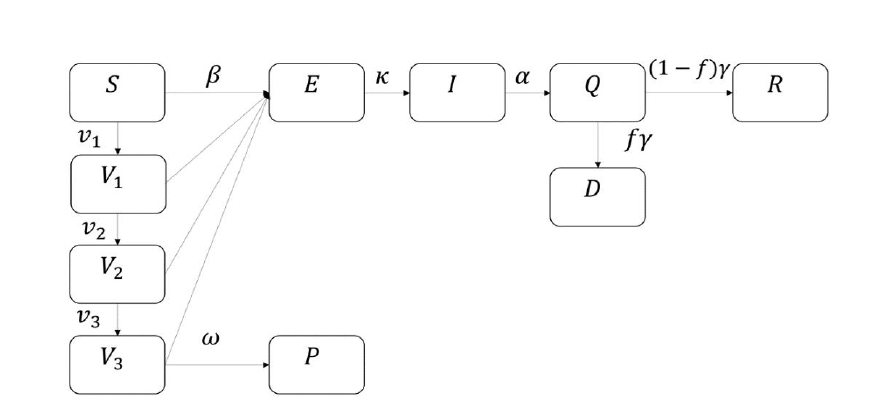
Fig.┬Ā3.
Fitted and predicted daily casesŌĆÖ curves using the SEIQRDV3P model for different Omicron transmission rates k = 1,3,5,7. The time axis combines both train and test data periods, ranging is from 20 September 2021 to 4 February 2022.

Table┬Ā1.
Previously determined model parameters
| Parameter | Description | Value |
|---|---|---|
| 1/╬║ | The average duration from E to I | 4.1 days [64-66] |
| 1/╬▒ | The average duration from I to Q | 6 days [64,65] |
| 1/╬│ | The average duration from Q to R or D | 20.1 days [67] |
| f | Mortality rate | 0.09 [68] |
| e | Efficacy of vaccination | 0.78 [69] |
References
2. Coronavirus disease (COVID-19) pandemic. Geneva: World Health Organization, 2020. Accessed 2022 Jan 20. Available from: https://www.who.int/emergencies/diseases/novel-coronavirus-2019.
3. Hsiang S, Allen D, Annan-Phan S, Bell K, Bolliger I, Chong T, et al. The effect of large-scale anti-contagion policies on the COVID-19 pandemic. Nature 2020;584:262ŌĆō267.



4. Ferguson NM, Laydon D, Nedjati-Gilani G, Imai N, Ainslie K, Baguelin M, et al. Impact of Non-pharmaceutical Interventions (NPIs) to Reduce COVID-19 Mortality and Healthcare Demand. London: Imperial College, 2020.
5. Brzezinski A, Deiana G, Kecht V, Van Dijcke D. The COVID-19 Pandemic: Government vs. Community Action across the United States. INET Oxford Working Paper No. 2020-06. Oxford: Institute for New Economic Thinking at the Oxford Martin School, University of Oxford, 2020.
8. Thanh Le T, Andreadakis Z, Kumar A, Gomez Roman R, Tollefsen S, Saville M, et al. The COVID-19 vaccine development landscape. Nat Rev Drug Discov 2020;19:305ŌĆō306.



9. Zuo M, Khosa SK, Ahmad Z, Almaspoor Z. Comparison of COVID-19 pandemic dynamics in Asian countries with statistical modeling. Comput Math Methods Med 2020;2020:4296806.




10. Muse AH, Tolba AH, Fayad E, Abu Ali OA, Nagy M, Yusuf M. Modelling the COVID-19 mortality rate with a new versatile modification of the log-logistic distribution. Comput Intell Neurosci 2021;2021:8640794.




11. de la Fuente-Mella H, Rubilar R, Chahuan-Jimenez K, Leiva V. Modeling COVID-19 cases statistically and evaluating their effect on the economy of countries. Mathematics 2021;9:1558.

12. Liu X, Ahmad Z, Gemeay AM, Abdulrahman AT, Hafez EH, Khalil N. Modeling the survival times of the COVID-19 patients with a new statistical model: a case study from China. PLoS One 2021;16:e0254999.



13. Biggerstaff M, Cowling BJ, Cucunuba ZM, Dinh L, Ferguson NM, Gao H, et al. Early insights from statistical and mathematical modeling of key epidemiologic parameters of COVID-19. Emerg Infect Dis 2020;26:e1ŌĆōe14.


14. Childs ML, Kain MP, Harris MJ, Kirk D, Couper L, Nova N, et al. The impact of long-term non-pharmaceutical interventions on COVID-19 epidemic dynamics and control: the value and limitations of early models. Proc Biol Sci 2021;288:20210811.




15. Moreau VH. Forecast predictions for the COVID-19 pandemic in Brazil by statistical modeling using the Weibull distribution for daily new cases and deaths. Braz J Microbiol 2020;51:1109ŌĆō1115.




16. Datta R, Trivedi PK, Kumawat A, Kumar R, Bhardwaj I, Kumari N, et al. Statistical modeling of COVID-19 pandemic stages worldwide. Preprints at https://doi.org/10.20944/preprints202005.0319.v1 (2020).

17. Roy S, Bhunia GS, Shit PK. Spatial prediction of COVID-19 epidemic using ARIMA techniques in India. Model Earth Syst Environ 2021;7:1385ŌĆō1391.



18. Singh RK, Rani M, Bhagavathula AS, Sah R, Rodriguez-Morales AJ, Kalita H, et al. Prediction of the COVID-19 pandemic for the top 15 affected countries: advanced autoregressive integrated moving average (ARIMA) model. JMIR Public Health Surveill 2020;6:e19115.



19. Zeroual A, Harrou F, Dairi A, Sun Y. Deep learning methods for forecasting COVID-19 time-series data: a comparative study. Chaos Solitons Fractals 2020;140:110121.



20. Arora P, Kumar H, Panigrahi BK. Prediction and analysis of COVID-19 positive cases using deep learning models: a descriptive case study of India. Chaos Solitons Fractals 2020;139:110017.



21. Kapoor A, Ben X, Liu L, Perozzi B, Barnes M, Blais M, et al. Examining COVID-19 forecasting using spatio-temporal graph neural networks. Preprint at https://arxiv.org/abs/2007.03113 (2020).
22. Fritz C, Dorigatti E, Rugamer D. Combining graph neural networks and spatio-temporal disease models to predict COVID-19 cases in Germany. Sci Rep 2022;12:3930.


23. Rauf HT, Lali MI, Khan MA, Kadry S, Alolaiyan H, Razaq A, et al. Time series forecasting of COVID-19 transmission in Asia Pacific countries using deep neural networks. Pers Ubiquitous Comput 2021 Jan 10 [Epub]. https://doi.org/10.1007/s00779-020-01494-0.

24. Nabi KN, Tahmid MT, Rafi A, Kader ME, Haider MA. Forecasting COVID-19 cases: a comparative analysis between recurrent and convolutional neural networks. Results Phys 2021;24:104137.



25. Chen TM, Rui J, Wang QP, Zhao ZY, Cui JA, Yin L. A mathematical model for simulating the phase-based transmissibility of a novel coronavirus. Infect Dis Poverty 2020;9:24.




26. Ndairou F, Area I, Nieto JJ, Torres DF. Mathematical modeling of COVID-19 transmission dynamics with a case study of Wuhan. Chaos Solitons Fractals 2020;135:109846.



27. Car Z, Baressi Segota S, Andelic N, Lorencin I, Mrzljak V. Modeling the spread of COVID-19 infection using a multilayer perceptron. Comput Math Methods Med 2020;2020:5714714.




28. Panovska-Griffiths J. Can mathematical modelling solve the current Covid-19 crisis? BMC Public Health 2020;20:551.




29. Kermack WO, McKendrick AG. A contribution to the mathematical theory of epidemics. Proc R Soc Lond Ser A Contain Pap Math Phys Character 1927;115:700ŌĆō721.


30. Shankar S, Mohakuda SS, Kumar A, Nazneen PS, Yadav AK, Chatterjee K, et al. Systematic review of predictive mathematical models of COVID-19 epidemic. Med J Armed Forces India 2021;77(Suppl 2):S385ŌĆōS392.



31. Kucharski AJ, Russell TW, Diamond C, Liu Y, Edmunds J, Funk S, et al. Early dynamics of transmission and control of COVID-19: a mathematical modelling study. Lancet Infect Dis 2020;20:553ŌĆō558.



32. Mohd MH, Sulayman F. Unravelling the myths of R 0 in controlling the dynamics of COVID-19 outbreak: a modelling perspective. Chaos Solitons Fractals 2020;138:109943.



33. Volpert V, Banerjee M, Petrovskii S. On a quarantine model of coronavirus infection and data analysis. Math Model Nat Phenom 2020;15:24.

34. Kochanczyk M, Grabowski F, Lipniacki T. Dynamics of COVID-19 pandemic at constant and time-dependent contact rates. Math Model Nat Phenom 2020;15:28.

35. Roda WC, Varughese MB, Han D, Li MY. Why is it difficult to accurately predict the COVID-19 epidemic? Infect Dis Model 2020;5:271ŌĆō281.



36. Tuite AR, Fisman DN, Greer AL. Mathematical modelling of COVID-19 transmission and mitigation strategies in the population of Ontario, Canada. CMAJ 2020;192:E497ŌĆōE505.



37. Hellewell J, Abbott S, Gimma A, Bosse NI, Jarvis CI, Russell TW, et al. Feasibility of controlling COVID-19 outbreaks by isolation of cases and contacts. Lancet Glob Health 2020;8:e488ŌĆōe496.



38. Liu Y, Gayle AA, Wilder-Smith A, Rocklov J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J Travel Med 2020;27:taaa021.



39. Cakir Z, Savas HB. A mathematical modelling approach in the spread of the novel 2019 coronavirus SARS-CoV-2 (COVID-19) pandemic. Electron J Gen Med 2020;17:em205.

40. Bouchnita A, Jebrane A. A multi-scale model quantifies the impact of limited movement of the population and mandatory wearing of face masks in containing the COVID-19 epidemic in Morocco. Math Model Nat Phenom 2020;15:31.

41. Sahin U, Sahin T. Forecasting the cumulative number of confirmed cases of COVID-19 in Italy, UK and USA using fractional nonlinear grey Bernoulli model. Chaos Solitons Fractals 2020;138:109948.



42. Roosa K, Lee Y, Luo R, Kirpich A, Rothenberg R, Hyman JM, et al. Real-time forecasts of the COVID-19 epidemic in China from February 5th to February 24th, 2020. Infect Dis Model 2020;5:256ŌĆō263.



43. Castro MC, Singer B. Prioritizing COVID-19 vaccination by age. Proc Natl Acad Sci U S A 2021;118:e2103700118.



44. Ritchie H, Mathieu E, Rodes-Guirao L, Appel C, Giattino C, Ortiz-Ospina E, et al. Coronavirus (COVID-19) vaccinations. Oxford: University of Oxford, 2020. Accessed 2022 Jan 22. Available from: https://ourworldindata.org/covid-vaccinations.
45. Vasireddy D, Vanaparthy R, Mohan G, Malayala SV, Atluri P. Review of COVID-19 variants and COVID-19 vaccine efficacy: what the clinician should know? J Clin Med Res 2021;13:317ŌĆō325.



47. Jo H, Son H, Hwang HJ, Jung SY. Analysis of COVID-19 spread in South Korea using the SIR model with time-dependent parameters and deep learning. Preprint at https://doi.org/10.1101/2020.04.13.20063412 (2020).

48. Kim YJ, Seo MH, Yeom HE. Estimating a breakpoint in the pattern of spread of COVID-19 in South Korea. Int J Infect Dis 2020;97:360ŌĆō364.



49. Nesteruk I. Estimations of the coronavirus epidemic dynamics in South Korea with the use of SIR model. Preprint at https://doi.org/10.13140/RG.2.2.15489.40807 (2020).

50. Cooper I, Mondal A, Antonopoulos CG. A SIR model assumption for the spread of COVID-19 in different communities. Chaos Solitons Fractals 2020;139:110057.



51. He J, Chen G, Jiang Y, Jin R, Shortridge A, Agusti S, et al. Comparative infection modeling and control of COVID-19 transmission patterns in China, South Korea, Italy and Iran. Sci Total Environ 2020;747:141447.



52. Reis RF, de Melo Quintela B, de Oliveira Campos J, Gomes JM, Rocha BM, Lobosco M, et al. Characterization of the COVID-19 pandemic and the impact of uncertainties, mitigation strategies, and underreporting of cases in South Korea, Italy, and Brazil. Chaos Solitons Fractals 2020;136:109888.



53. Amiri Mehra AH, Shafieirad M, Abbasi Z, Zamani I. Parameter estimation and prediction of COVID-19 epidemic turning point and ending time of a case study on SIR/SQAIR epidemic models. Comput Math Methods Med 2020;2020:1465923.


54. Ko Y, Lee J, Seo Y, Jung E. Risk of COVID-19 transmission in heterogeneous age groups and effective vaccination strategy in Korea: a mathematical modeling study. Epidemiol Health 2021;43:e2021059.



55. Domingo E, Holland JJ. RNA virus mutations and fitness for survival. Annu Rev Microbiol 1997;51:151ŌĆō178.


56. Zhou W, Wang W. Fast-spreading SARS-CoV-2 variants: challenges to and new design strategies of COVID-19 vaccines. Signal Transduct Target Ther 2021;6:226.




57. Ko Y, Lee J, Kim Y, Kwon D, Jung E. COVID-19 vaccine priority strategy using a heterogenous transmission model based on maximum likelihood estimation in the Republic of Korea. Int J Environ Res Public Health 2021;18:6469.



58. Cooper I, Mondal A, Antonopoulos CG. A SIR model assumption for the spread of COVID-19 in different communities. Chaos Solitons Fractals 2020;139:110057.



59. Kim S, Seo YB, Jung E. Prediction of COVID-19 transmission dynamics using a mathematical model considering behavior changes in Korea. Epidemiol Health 2020;42:e2020026.


60. Brauer F. Compartmental models in epidemiology. In: Mathematical Epidemiology (Brauer F, van den Driessche P, Wu J, eds.). Berlin: Springer Berlin Heidelberg, 2008. pp. 19ŌĆō79.
61. JM24. fitVirusCV19v3 (COVID-19 SIR Model). Natick: MathWorks, 2022;Accessed 2022 Jan 23. Available from: https://www.mathworks.com/matlabcentral/fileexchange/74676-fitviruscv19v3-covid-19-sir-model.
62. Hale T, Petherick A, Phillips T, Webster S. Variation in Government Responses to COVID-19. Blavatnik School of Government Working Paper 31. Oxford: Blavatnik School of Government, 2020.
63. Blavatnik School of Government, University of Oxford. COVID-19 government esponse tracker. Oxford: Blavatnik School of Government, 2020. Accessed 2022 Jan 22. Available from: https://www.bsg.ox.ac.uk/research/research-projects/coronavirus-government-response-tracker.
64. Korean situation report of COVID-19. Cheonju: Korea Disease Control and Prevention Agency, 2020, Accessed 2022 Jan 4. Available from: http://ncov.mohw.go.kr/tcmBoardView.do?brdId=&brdGubun=&dataGubun=&ncvContSeq=352903&contSeq=352903.
65. Ganyani T, Kremer C, Chen D, Torneri A, Faes C, Wallinga J, et al. Estimating the generation interval for coronavirus disease (COVID-19) based on symptom onset data, March 2020. Euro Surveill 2020;25:2000257.



66. Ki M, Task Force for 2019-nCoV. Epidemiologic characteristics of early cases with 2019 novel coronavirus (2019-nCoV) disease in Korea. Epidemiol Health 2020;42:e2020007.



67. Lee YH, Hong CM, Kim DH, Lee TH, Lee J. Clinical course of asymptomatic and mildly symptomatic patients with coronavirus disease admitted to community treatment centers, South Korea. Emerg Infect Dis 2020;26:2346ŌĆō2352.



68. Ritchie H, Mathieu E, Rodes-Guirao L, Appel C, Giattino C, Ortiz-Ospina E, et al. Coronavirus pandemic (COVID-19). Oxford: University of Oxford, 2020. Accessed 2022 Jan 4. Available from: https://ourworldindata.org/coronavirus.
69. Quick review on COVID-19 vaccine issues (ver. 2.0). Seoul: National Evidence-based healthcare Collaborating Agency, 2021. Available from: https://www.neca.re.kr/lay1/bbs/S1T11C174/F/58/view.do?article_seq=8634.
70. Mathieu E, Ritchie H, Ortiz-Ospina E, Roser M, Hasell J, Appel C, et al. A global database of COVID-19 vaccinations. Nat Hum Behav 2021;5:947ŌĆō953.



71. Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis 2020;20:533ŌĆō534.



72. Elbe S, Buckland-Merrett G. Data, disease and diplomacy: GISAID's innovative contribution to global health. Glob Chall 2017;1:33ŌĆō46.




73. Shu Y, McCauley J. GISAID: Global initiative on sharing all influenza data: from vision to reality. Euro Surveill 2017;22:30494.



74. Living with COVID-19: what to expect. Seoul: Korean Herald, 2021. Accessed 2022 Feb 11. Available from: http://www.koreaherald.com/view.php?ud=20211102000490.
- TOOLS
-
METRICS

-
- 3 Crossref
- 0 Scopus
- 5,424 View
- 190 Download









