 |
 |
- Search
| Genomics Inform > Volume 17(4); 2019 > Article |
|
Abstract
The incidence and mortality rate of cancer continues to gradually increase, although considerable research effort has been directed at elucidating the molecular mechanisms underlying biomarkers responsible for tumorigenesis. Accumulated evidence indicates that the long non-coding RNAs (lncRNAs), which are transcribed but not translated into functional proteins, contribute to cancer development. Recently, linc00152 (an lncRNA) was identified as a potent oncogene in various cancer types, and shown to be involved in cancer cell proliferation, invasiveness, and motility by sponging tumor-suppressive microRNAs acting as a competing endogenous RNA, binding to gene promoters acting as a transcriptional regulator, and binding to functional proteins. In this review, we focus on the oncogenic role of linc00152 in tumorigenesis and provided an overview of recent clinical studies on the effects of linc00152 expression in human cancers.
Cancers are diseases caused by abnormal cell growth. Unlike benign tumors where cells grow abnormally but do not spread, malignant tumors can migrate to other parts of the body. Cancers occur when genes are mutated and cell growth is abnormally regulated. Currently, oncogenes that cause cell cycle progression and contribute to cell proliferation, and tumor suppressor genes that participate in cell cycle arrest are subjects of active research. Typical examples are ERBB2 encoding oncogene HER2/neu [1], and TP53 encoding tumor suppressor gene p53 [2]. Recent studies have reported that genes not encoding a specific protein, such as long non-coding RNAs (lncRNAs) also affect the development of cancer.
LncRNAs are RNA transcripts that are longer than 200 nucleotides and are not translated into proteins. Many efforts have been made to identify the biological roles of lncRNAs in nuclei and cytoplasm. In nuclei, lncRNAs recruit chromatin-modifying enzymes and act as transcriptional guides. LncRNAs can also bind to mRNA and splicing factors and participate in splicing processes [3,4]. Other lncRNAs can bind to promoters of specific genes and facilitate or suppress their transcription. In addition, lncRNAs may also target RNA polymerase and transcription factors and regulate the transcriptions and expressions of genes [5]. For example, lncRNA YY1 directly interacts with transcription factor YY1 in muscle cells to activate gene expression by removing YY1 and polycomb repressive complex (PRC2) from the promoters of target genes [6]. In cytoplasm, lncRNAs may serve as competing endogenous RNAs (ceRNAs) that target microRNAs (miRNAs) [5], and interactions between lncRNA and miRNA targets, disrupts binding between the miRNA and its target mRNA, and leads to the induction of mRNA translation. In addition, lncRNAs can interact with proteins possessing RNA binding motifs and increase their stability and activity [7,8]. Such lncRNAs participate in gene regulation at the transcriptional and translational levels and in protein activation at the post-translational levels. Recently, several studies have reported that based on their biological activities, lncRNAs are highly associated with various diseases including cancer [9,10]. When cancer cells with counterpart normal cells were compared in various types of cancer, abnormal lncRNA expressions were observed in cancer cells. In addition, lncRNAs were found to regulate cell growth and proliferation during cancer development. Moreover, it was recently reported that long non-coding RNA linc00641 can sponge miR-424-5p in non-small cell lung cancer cells and act as a tumor suppressor [11].
Linc00152 (also known as cytoskeleton regulator RNA [CYTOR]) is an lncRNA located at 2p 11.2. Studies have shown that linc00152 is overexpressed in cancer cells and promotes cancer cell proliferation and metastasis. For example, it was found that the upregulation of linc00152 is associated with enhanced cell invasion in gastric adenocarcinoma cells [12]. In general, linc00152 is involved in the sponging of miRNAs or in the transcriptional silencing of tumor suppressor genes; activities that lead to cell proliferation and epithelial-mesenchymal transition. Here, we focus on the role of linc00152 during cancer development and summarize the findings of recent studies on the effects of linc00152 expression on cancer progression and on the molecular mechanisms of linc00152 in human cancers.
One of the roles of linc00152 is to function as a ceRNA of miRNA. The ceRNA activity of linc00152 may be associated with the upregulations of various counterpart mRNAs (originally targeted by miRNAs) involved in cell proliferation, survival, protection from apoptosis, invasion, and migration (Table 1). It has been reported that linc00152 might directly bind to miR-125b, up-regulate Mcl-1 (myeloid cell leukemia-1), and protect ovarian adenocarcinoma cells from apoptosis [13]. Linc00152 might also sponge miR-193a-3p, and thus, contribute to the upregulation of Mcl-1 expression [14]. In addition, linc00152 may promote cell proliferation and increase cell migration in gastric adenocarcinoma cells by sponging miR-193b-3p, and thus, upregulating ETS1 [15]. In another study, it was found that sponging of miR-139-5p by linc00152 up-regulated Notch1, which promoted cell proliferation, cell invasion, and migration in colorectal carcinoma cells [16]. Linc00152 might also promote cell proliferation by sponging miR-216b-5p, and thus, up-regulate the expression of homeobox A1 [17]. In addition, linc00152 might enhance the invasion and migration capacities of cancer cells by sponging miR-138 and upregulating hypoxia-inducible factor-1α (HIF-1α) expression [18]. Linc00152 may also interact directly with miR-139-5p, which is associated with the expressional upregulation of protein kinase AMP-activated catalytic subunit alpha 1, which is involved in the promotion of aerobic glycolysis for metabolic reprogramming [19]. These results suggest the associations between linc00152 and HIF-1α activation and increased aerobic glycolysis might be linked to cancer cell survival against intratumoral hypoxia, and contribute to tumor aggressiveness and malignant development. Several studies have reported that linc00152 might be involved in cell cycle regulation through its miRNA-sponging activities. In one study conducted using osteosarcoma cells, it was suggested that linc00152 transcriptionally activated by TCF3 (a transcription factor) might bind to miR-1182, up-regulate CDK14, and promote cell proliferation and migration [20]. In another study, linc00152 was found to up-regulate cyclin D1 and cyclin-dependent kinase 9 by sponging miR-193a [21,22]. Furthermore, it was reported that linc00152 increased the expressions of several proto-oncogenes by sponging their counterpart miRNAs in various types of cancer. It was suggested that linc00152 might down-regulate miR-612, be involved in the overexpression of Akt2, contribute to the activation of the nuclear factor kappa-light chain-enhancer of activated B cells (NF-κB) pathway, and consequently inhibit glioblastoma cell apoptosis [23]. In another study, results indicated that linc00152-dependent miR-153-3p down-regulation up-regulated Fyn (a proto-oncogene) and led to the induction of cell proliferation and the suppression of apoptosis in esophageal squamous cell carcinoma cells [24]. Thus, many studies have shown that linc00152 might be responsible for upregulations of several genes by direct binding miRNAs and disrupting their interactions with target mRNAs. Consequently, gene products up-regulated by the ceRNA activity of linc00152 may be involved in the promotions of cell proliferation, survival, and cell motility and in the suppression of apoptosis, which suggests linc00152 is a potential oncogenic ceRNA that promotes cancer progression.
Linc00152 can bind to the promoter regions of specific genes and regulate their transcriptions (Fig. 1). One study presented that linc00152 can bind to the promoter of epithelial adhesion molecule (EpCAM) gene, and thus, induce cell proliferation and tumor growth by inducing the transcriptional upregulation of EpCAM and activation of the mammalian target of rapamycin pathway [25]. In addition, linc00152 may participate in the transcriptional repression of interleukin 24 (IL24) caused by the recruitment of enhancer of zeste homolog 2 (EZH2) to IL24 promoter. EZH2 as a histone methyltransferase participate in trimethylation of histone H3 Lys 27 at the IL24 promoter region, and thereby, facilitated lung adenocarcinoma cell growth [26]. Linc00152 also recruited EZH2 to p15 and p21 promoters, and thus, inhibited the transcriptions of p15 and p21, which increased cell cycle progression and cell proliferation [27]. In addition, linc00152 caused the proliferation of bladder carcinoma cells by upregulating β-catenin expression and directly activating the Wnt/β-catenin pathway [28]. Several studies on the roles of linc00152 in cancer cells have reported that linc00152 was involved in the transcriptional activations of oncogenes by directly binding to gene promoters and in the transcriptional repressions of tumor suppressors by interacting with other transcription factors. These findings indicate that linc00152 functions as an oncogenic transcriptional regulator.
In addition to the involvement of linc00152 in transcription regulation via binding to the promoter regions of genes or miRNAs, linc00152 can also promote cell proliferation by directly binding to specific proteins and regulating their activities (Fig. 2). One study showed that linc00152 is capable of complex formation with two RNA binding proteins, NCL (nucleolin) and Sam68 (KHDRBS1, the src-associated substrate in mitosis of 68kDa), which are responsible for the development of colorectal cancer via NF-κB pathway activation [29]. Another study presented that linc00152 directly bound to epidermal growth factor receptor (EGFR) and increased the activity of EGFR to promote the EGFR/phosphoinositide 3-kinase/Akt pathway, which contributed to the inductions of tumorigenic features (e.g., cell cycle progression, cell proliferation, and migration) [30,31]. Moreover, linc00152 might be involved in the ubiquitin-dependent degradations of phosphatase and tensin homolog (PTEN) through the activation of NEDD4-1 (neural precursor cell expressed developmentally down-regulated protein 4-1) in breast adenocarcinoma [32]. It has also been reported that interaction between linc00152 and DNA methyltransferase (DNMT) may result in DNMT activation, the inhibitions of BRCA1 (breast cancer type 1 susceptibility protein) and PTEN, and increased cell proliferation and invasiveness in triple-negative breast adenocarcinoma and breast ductal carcinoma [33]. These studies show that proteins affected by linc00152 are involved in signaling pathways that contribute to cell survival, proliferation, and apoptosis, and suggest that linc00152 might facilitate malignant progression by activating cell survival/proliferation pathways and inhibiting of apoptotic pathways by interacting with specific proteins and by upregulating the transcriptional expressions of oncogenes.
Several studies have addressed relations between linc00152 expression and malignant development. These studies revealed that linc00152 was overexpressed in various cancer types, including liver cancer, gastric cancer, lung cancer, and breast cancer [14,24,30,33]. Moreover, clinical studies have shown that high linc00152 expression in tumor tissues is associated with poor survival and disease-free survival [17,21,23,29,34]. Linc00152 expression has also been shown to be significantly correlated with tumor progression stage. Among cancer patients of TNM stage III or greater, the proportion of patients with high linc00152 expression was significantly higher than among patients with low linc00152 expression [13,16,25-27,31,35]. In a study of esophageal squamous cell carcinoma, patients with TNM stage I/II/III had higher linc00152 expressions [36]. Furthermore, linc00152 expression was also shown to be positively correlated with tumor size. Among patients with a tumor larger than 5 cm, the proportion of patients with high linc00152 expression was significantly greater than among patients with low expression [25-27,31,35], and in a study on osteosarcoma, a significantly greater proportion of patients with a tumor size of >3 cm exhibited ‘high’ linc00152 expression [20]. These results demonstrate that the expression of linc00152 is higher in tumor tissues than in normal tissues, that its expression is positively associated with tumor progression, and that high linc00152 expression is associated with poor prognosis in cancer patients. Furthermore, they indicate that linc00152 might act as a promising diagnostic and prognostic oncogene.
Many studies have investigated the association between linc00152 and human cancers. Linc00152 expression has been shown to be greater in tumor tissues than in normal tissues and to function as an oncogene by promoting cell proliferation, tumor growth, and metastasis in vivo. Linc00152 acts on cancer cells in three major ways: by acting as a ceRNA and sponging miRNAs, by binding to promoter regions of specific genes to activate or inhibit transcription, or by directly binding to and regulating the activities of proteins. In cancers (e.g., ovarian, gastric, and colorectal cancer), linc00152 acts as a ceRNA and sponges various miRNAs, especially miR-125b, miR-193a, miR193b, miR-612, miR-138, miR-216b-5p, miR-153-3p, miR-1182 or miR-497, in cancer cells, and thus, increase the expressions of downstream genes targeted by miRNAs, and consequently, these increases promote cell proliferation, metastasis, and tumor development. Linc00152 might also participate in the transcriptional activations or repressions of specific genes (e.g., EpCAM, IL24, p15, and p21) and activate oncogenic signaling pathways by interacting with several proteins (e.g., EGFR and NCL/Sam68 complex). Accordingly, linc00152 facilitates cancer cell development by directly or indirectly controlling the expressions and activities of many genes involved in the cell cycle and cell proliferation. By integrating and analyzing the biological roles of linc00152 during cancer progression, linc00152 may prove to be a useful diagnostic and prognostic biomarker for human cancers.
Notes
Fig. 1.
Linc00152 can participate in gene transcription. Linc00152 bound to gene promoters and up-regulated expression of epithelial adhesion molecule (EpCAM) and β-catenine, contributing to cell proliferation and cancer development through the activation of mammalian target of rapamycin (mTOR) and Wnt/β-catenine pathways, respectively. In addition, linc00152 recruited enhancer of zeste homolog 2 (EZH2) to the promoter of p15, p21 and interleukin 24 (IL24) responsible for down-regulated expression of these genes, leading to promotion of cell proliferation and inhibition of apoptosis.
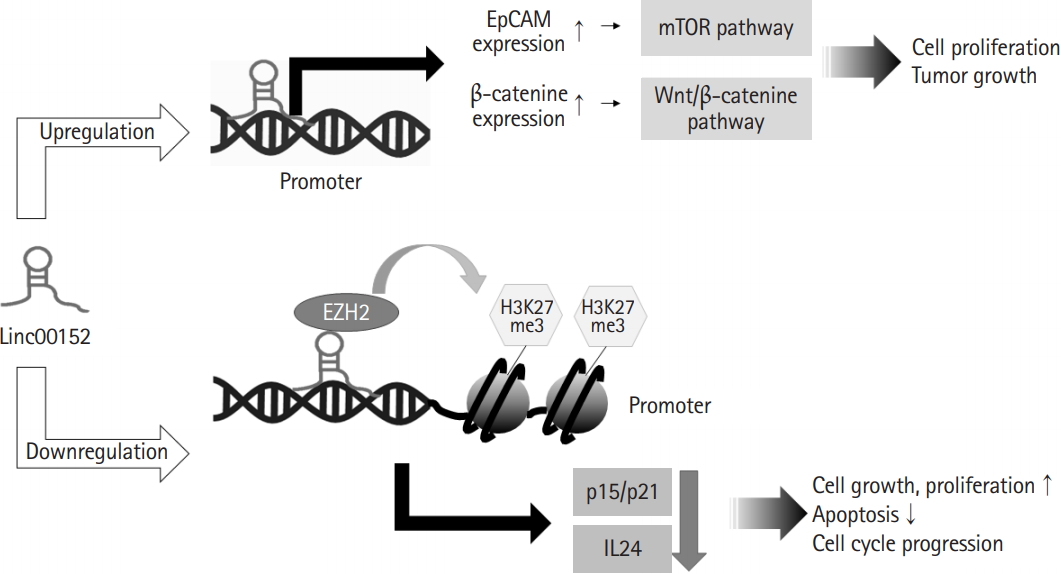
Fig. 2.
Linc00152 is involved in protein activation. Linc00152 bound directly to epidermal growth factor receptor (EGFR). The activated EGFR/phosphoinositide 3-kinase (PI3K)/AKT pathway promoted cell proliferation and migration. In addition, the complex of linc00152 and Sam68 promoted tumor progression through the activation of nuclear factor kappa-light chain-enhancer of activated B cells (NF-κB) pathway. DNA methyltransferase (DNMT) and NEDD4-1 (neural precursor cell expressed developmentally down-regulated protein 4-1) activated by linc00152 promoted cancer cell proliferation through inhibition of breast cancer type1 susceptibility protein (BRCA1) and phosphatase and tensin homolog (PTEN).
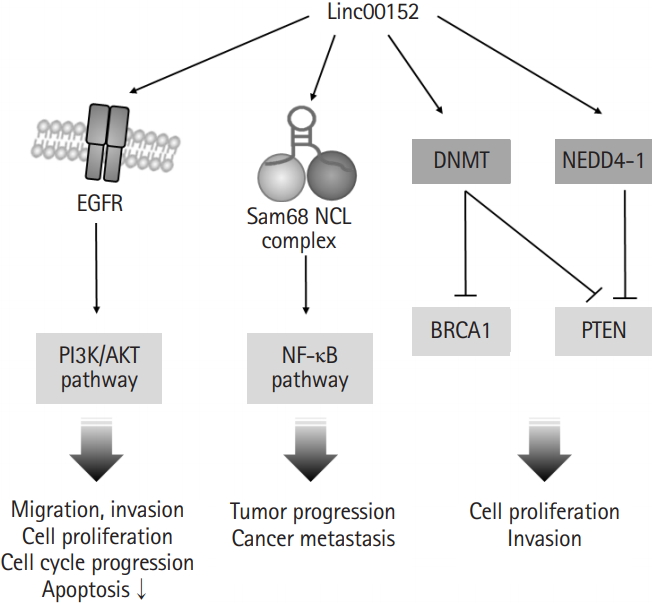
Table 1.
Linc00152-sponged miRNAs and their target genes, biological consequences in cancer cells
| Target miRNA | Target gene of miRNA | Biological consequences | Cancer type | Reference |
|---|---|---|---|---|
| miR-125b | Mcl-1 | Inhibition of apoptosis | Ovarian adenocarcinoma | [13] |
| miR-193a-3p | Cell proliferation | Gastric adenocarcinoma | [14] | |
| miR-193b-3p | ETS1 | Cell proliferation and migration | Gastric adenocarcinoma | [15] |
| miR-139-5p | PRKAA1 | Glycolytic phenotype | Gastric adenocarcinoma | [19] |
| Notch1 | Cell proliferation, cell invasion and migration | Colorectal carcinoma | [16] | |
| miR-193a | CDK9 | Cell cycle progression, cell proliferation, and inhibition of apoptosis | Acute monocytic leukemia, acute promyelocytic leukemia | [21] |
| miR-612 | Akt2 | Tumor growth, cell invasion, and inhibition of apoptosis | Glioblastoma | [23] |
| miR-138 | HIF-1α | Cell invasion and migration | Gallbladder carcinoma | [18] |
| miR-216b-5p | HOXA1 | Cell proliferation | Cervix epidermoid carcinoma | [17] |
| miR-153-3p | Fyn | Cell proliferation and inhibition of apoptosis | Esophageal squamous cell carcinoma | [24] |
| miR-1182 | CDK14 | Cell proliferation and migration | Osteosarcoma | [20] |
References
1. Gschwind A, Fischer OM, Ullrich A. The discovery of receptor tyrosine kinases: targets for cancer therapy. Nat Rev Cancer 2004;4:361–370.



2. Surget S, Khoury MP, Bourdon JC. Uncovering the role of p53 splice variants in human malignancy: a clinical perspective. Onco Targets Ther 2013;7:57–68.


3. Nobili L, Ronchetti D, Taiana E, Neri A. Long non-coding RNAs in B-cell malignancies: a comprehensive overview. Oncotarget 2017;8:60605–60623.



4. Do H, Kim W. Roles of oncogenic long non-coding RNAs in cancer development. Genomics Inform 2018;16:e18.




5. Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol 2018;19:143–157.



6. Zhou L, Sun K, Zhao Y, Zhang S, Wang X, Li Y, et al. Linc-YY1 promotes myogenic differentiation and muscle regeneration through an interaction with the transcription factor YY1. Nat Commun 2015;6:10026.



7. Paraskevopoulou MD, Hatzigeorgiou AG. Analyzing miRNA-lncRNA interactions. Methods Mol Biol 2016;1402:271–286.


8. Zhu J, Fu H, Wu Y, Zheng X. Function of lncRNAs and approaches to lncRNA-protein interactions. Sci China Life Sci 2013;56:876–885.



9. Ma L, Cao J, Liu L, Du Q, Li Z, Zou D, et al. LncBook: a curated knowledgebase of human long non-coding RNAs. Nucleic Acids Res 2019;47:D128–D134.



10. Chen X, Yan CC, Zhang X, You ZH. Long non-coding RNAs and complex diseases: from experimental results to computational models. Brief Bioinform 2017;18:558–576.



11. Li Y, Zhao L, Zhao P, Liu Z. Long non-coding RNA LINC00641 suppresses non-small-cell lung cancer by sponging miR-424-5p to upregulate PLSCR4. Cancer Biomark 2019;26:79–91.


12. Pang Q, Ge J, Shao Y, Sun W, Song H, Xia T, et al. Increased expression of long intergenic non-coding RNA LINC00152 in gastric cancer and its clinical significance. Tumour Biol 2014;35:5441–5447.



13. Chen P, Fang X, Xia B, Zhao Y, Li Q, Wu X. Long noncoding RNA LINC00152 promotes cell proliferation through competitively binding endogenous miR-125b with MCL-1 by regulating mitochondrial apoptosis pathways in ovarian cancer. Cancer Med 2018;7:4530–4541.



14. Huang Y, Luo H, Li F, Yang Y, Ou G, Ye X, et al. LINC00152 down-regulated miR-193a-3p to enhance MCL1 expression and promote gastric cancer cells proliferation. Biosci Rep 2018;38:BSR20171607.




15. Wang H, Chen W, Yang P, Zhou J, Wang K, Tao Q. Knockdown of linc00152 inhibits the progression of gastric cancer by regulating microRNA-193b-3p/ETS1 axis. Cancer Biol Ther 2019;20:461–473.


16. Bian Z, Zhang J, Li M, Feng Y, Yao S, Song M, et al. Long non-coding RNA LINC00152 promotes cell proliferation, metastasis, and confers 5-FU resistance in colorectal cancer by inhibiting miR-139-5p. Oncogenesis 2017;6:395.



17. Zheng JJ, Du XJ, Wang HP, Zhou LY, Wang YJ, Zhang L, et al. Long non-coding RNA 00152 promotes cell proliferation in cervical cancer via regulating miR-216b-5p/HOXA1 axis. Eur Rev Med Pharmacol Sci 2019;23:3654–3663.

18. Cai Q, Wang Z, Wang S, Weng M, Zhou D, Li C, et al. Long non-coding RNA LINC00152 promotes gallbladder cancer metastasis and epithelial-mesenchymal transition by regulating HIF-1α via miR-138. Open Biol 2017;7:160247.



19. Sun K, Hu P, Xu F. LINC00152/miR-139-5p regulates gastric cancer cell aerobic glycolysis by targeting PRKAA1. Biomed Pharmacother 2018;97:1296–1302.


20. Zheng L, Hu N, Zhou X. TCF3-activated LINC00152 exerts oncogenic role in osteosarcoma through regulating miR-1182/CDK14 axis. Pathol Res Pract 2019;215:373–380.


21. Ma P, Wang H, Sun J, Liu H, Zheng C, Zhou X, et al. LINC00152 promotes cell cycle progression in hepatocellular carcinoma via miR-193a/b-3p/CCND1 axis. Cell Cycle 2018;17:974–984.



22. Zhang X, Tao W. Long noncoding RNA LINC00152 facilitates the leukemogenesis of acute myeloid leukemia by promoting CDK9 through miR-193a. DNA Cell Biol 2019;38:236–242.


23. Cai J, Zhang J, Wu P, Yang W, Ye Q, Chen Q, et al. Blocking LINC00152 suppresses glioblastoma malignancy by impairing mesenchymal phenotype through the miR-612/AKT2/NF-κB pathway. J Neurooncol 2018;140:225–236.



24. Liu D, Gao M, Wu K, Zhu D, Yang Y, Zhao S. LINC00152 facilitates tumorigenesis in esophageal squamous cell carcinoma via miR-153-3p/FYN axis. Biomed Pharmacother 2019;112:108654.


25. Ji J, Tang J, Deng L, Xie Y, Jiang R, Li G, et al. LINC00152 promotes proliferation in hepatocellular carcinoma by targeting EpCAM via the mTOR signaling pathway. Oncotarget 2015;6:42813–42824.



26. Chen QN, Chen X, Chen ZY, Nie FQ, Wei CC, Ma HW, et al. Long intergenic non-coding RNA 00152 promotes lung adenocarcinoma proliferation via interacting with EZH2 and repressing IL24 expression. Mol Cancer 2017;16:17.




27. Chen WM, Huang MD, Sun DP, Kong R, Xu TP, Xia R, et al. Long intergenic non-coding RNA 00152 promotes tumor cell cycle progression by binding to EZH2 and repressing p15 and p21 in gastric cancer. Oncotarget 2016;7:9773–9787.



28. Xian-Li T, Hong L, Hong Z, Yuan L, Jun-Yong D, Peng X, et al. Higher expression of Linc00152 promotes bladder cancer proliferation and metastasis by activating the Wnt/β-catenin signaling pathway. Med Sci Monit 2019;25:3221–3230.



29. Wang X, Yu H, Sun W, Kong J, Zhang L, Tang J, et al. The long non-coding RNA CYTOR drives colorectal cancer progression by interacting with NCL and Sam68. Mol Cancer 2018;17:110.




30. Zhang Y, Xiang C, Wang Y, Duan Y, Liu C, Jin Y, et al. lncRNA LINC00152 knockdown had effects to suppress biological activity of lung cancer via EGFR/PI3K/AKT pathway. Biomed Pharmacother 2017;94:644–651.


31. Zhou J, Zhi X, Wang L, Wang W, Li Z, Tang J, et al. Linc00152 promotes proliferation in gastric cancer through the EGFR-dependent pathway. J Exp Clin Cancer Res 2015;34:135.



32. Shen X, Zhong J, Yu P, Zhao Q, Huang T. YY1-regulated LINC00152 promotes triple negative breast cancer progression by affecting on stability of PTEN protein. Biochem Biophys Res Commun 2019;509:448–454.


33. Wu J, Shuang Z, Zhao J, Tang H, Liu P, Zhang L, et al. Linc00152 promotes tumorigenesis by regulating DNMTs in triple-negative breast cancer. Biomed Pharmacother 2018;97:1275–1281.


34. Yu T, Xu Z, Zhang X, Men L, Nie H. Long intergenic non‑protein coding RNA 152 promotes multiple myeloma progression by negatively regulating microRNA‑497. Oncol Rep 2018;40:3763–3771.


- TOOLS
-
METRICS

-
- 13 Crossref
- 0 Scopus
- 5,736 View
- 94 Download
- Related articles in GNI







