Introduction
The human genome contains actively moving retrotransposons (human endogenous retrovirus [HERV], long interspersed nucleotide element [LINE], and short interspersed nucleotide element [SINE]). Approximately 36% of the human genome has been generated through retrotransposition of LINE and other RNA species by the LINE reverse transcriptase [1, 2]. Retrotransposition is progressing in the human population by L1, Alu, and SINE-VNTR-Alu (SVA) insertions [3]. L1 is an autonomous retrotransposon that contains an internal RNA polymerase II promoter and a reverse transcriptase, whereas Alu and SVA lack activities for independent mobilization [4, 5]. Therefore, Alu and SVA are assumed to use the L1 protein machinery for their own mobilization [3], and retrotransposition events of marked SVA elements occur indeed by L1 elements in human cultured cells [5, 6].
The SVA elements were originally named from the SINE-R retroposon, derived from an endogenous retrovirus, the HERV-K LTR element. SINE-R11, 14, and 19 have been isolated by colony blot hybridization using the LTR element as probe [7]. SINE-R.C2 has been found in the third intron of the C2 gene on the short arm of human chromosome 6, which was a human-specific element [8]. Within the Xq21.3 block, two SINE-R retroposons (HS307 and HS408) were identified [9]. Multiple copy numbers of retroposons have been successively detected in hominoid primates and humans [9-16]. Other similar sequences have been associated with Alu-like sequences within the HLA-RP1 (STK19) gene [17]. These composite retroposons with the entire structure are named SVA (SINE-R, VNTR, and Alu).
The SVA families are evolutionarily young and hominoid-specific retroelements. They have ability to influence a genomic locus in which they reside and cause various human diseases as insertional mutagens. In fact, Fukuyama muscular dystrophy (FCMD) is derived from the polymorphic SVA-E insertion in the 3' untranslated region (UTR) of the FUKUTIN gene [18]. SVA elements could also drive transcription of functional genes. In the 5' upstream region of TBPL2, the SVA element serves as a promoter, while the SVA-D element in the WDR66 gene promotes the transcription of a human-specific transcript variant [19]. Here, we analyzed structure variants of functional genes mediated by SVA subfamilies and examined their expression patterns in various human tissues.
Methods
Bioinformatic analysis
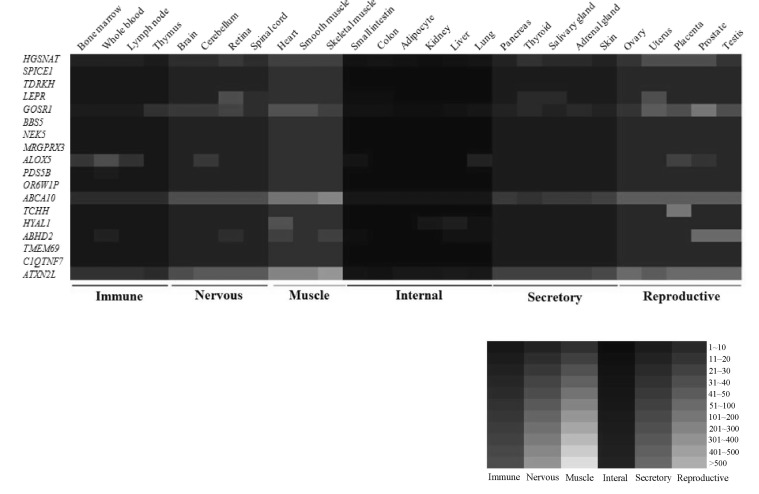
To identify SVA consensus sequences in the human genome, we obtained SVA sequences from the Giri database (http://www.girinst.org). The SVA subfamilies were aligned using the BioEdit program [20]. Then, we identified SVAs in each region. RepeatMasker (http://www.repeatmasker.org) and the UCSC genome site (http://genome.ucsc.edu) were employed to analyze isoform structures of functional genes. The human expressed sequence tag and RefSeq mRNA were also used to identify alternatively spliced transcripts. The expression pattern of SVA fusion genes in normal human tissues was analyzed using GeneCard (http://www.genecards.org). We obtained microarray data from the BioGPS database, and then we generated a heatmap according to microarray values. High expression levels were indicated by brighter color, and low expression levels were indicated by darker color.
Human RNA samples
A human 20-RNA tissue master panel (1, adrenal gland; 2, bone marrow; 3, cerebellum; 4, whole brain; 5, fetal brain; 6, fetal liver; 7, heart; 8, kidney; 9, liver; 10, lung; 11, placenta; 12, prostate; 13, salivary gland; 14, skeletal muscle; 15, spinal cord; 16, testis; 17, thymus; 18, thyroid; 19, trachea; 20, uterus) was purchased from Clontech (Mountain View, CA, USA).
Reverse-transcription (RT) and reverse transcription polymerase chain reaction (RT-PCR) amplification
To eliminate possible DNA contamination of purchased RNA samples, Turbo DNA-free (Ambion, Austin, TX, USA) was used according to the manufacturer's instructions. A no-RT control was also amplified to double-check the absence of DNA contamination. Quantity of RNA samples was measured using a ND-1000 UV-Vis spectrophotometer (NanoDrop, Wilmington, DE, USA). Moloney-Murine-Leukemia-Virus reverse transcriptase with an annealing temperature of 42℃ was used for the RT reaction with RNase inhibitor (Promega, Madison, WI, USA). To develop the specific primers for individual alternative transcripts, primer pairs were designed with the aid of Primer3 (http://frodo.wi.mit.edu/) (Table 1). In each run, 1 µL of cDNA was used as template for amplification per reaction. RT-PCR was performed using reactions containing a mixed cDNA template, representing a combination of different tissues examined. RT-PCR amplification for functional genes and a housekeeping gene was carried out for 30 cycles of 94℃ for 3 minutes, 56-60℃ for 1 minute, and 72℃ for 3 minutes. As a standard control, G3PDH was amplified through RT-PCR in human tissues. PCR products were loaded on 1-2% agarose gels and stained with ethidium bromide.
Results and Discussion
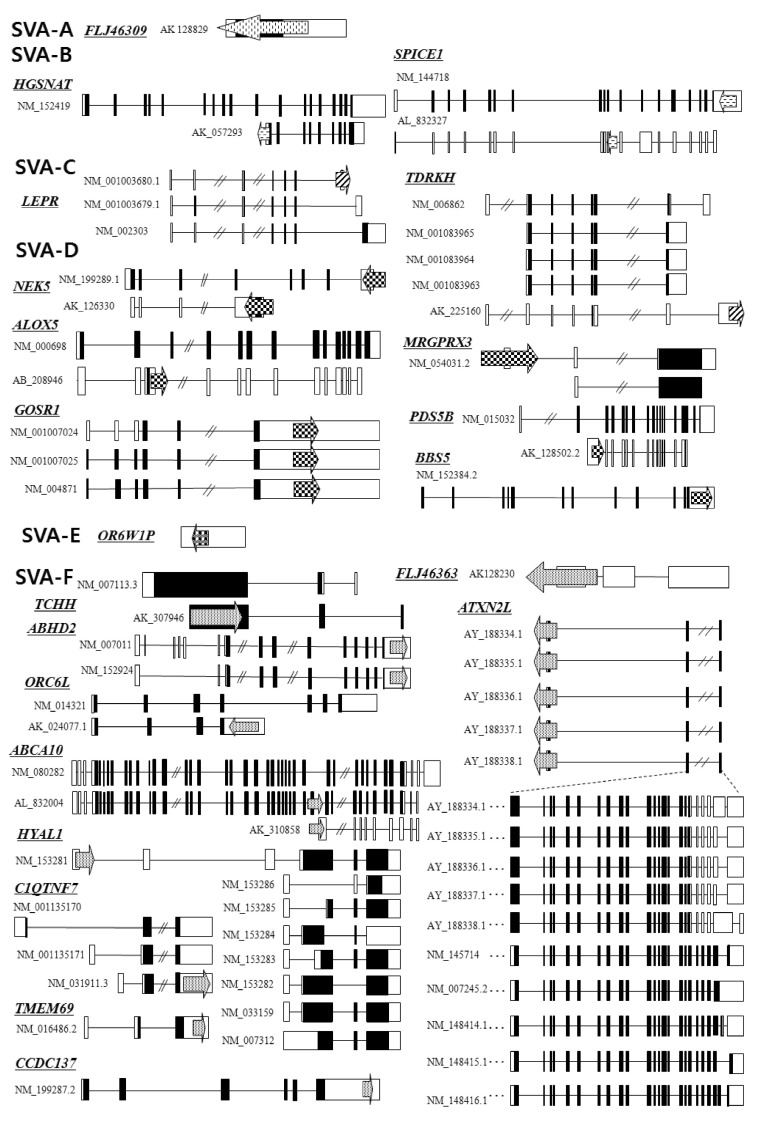
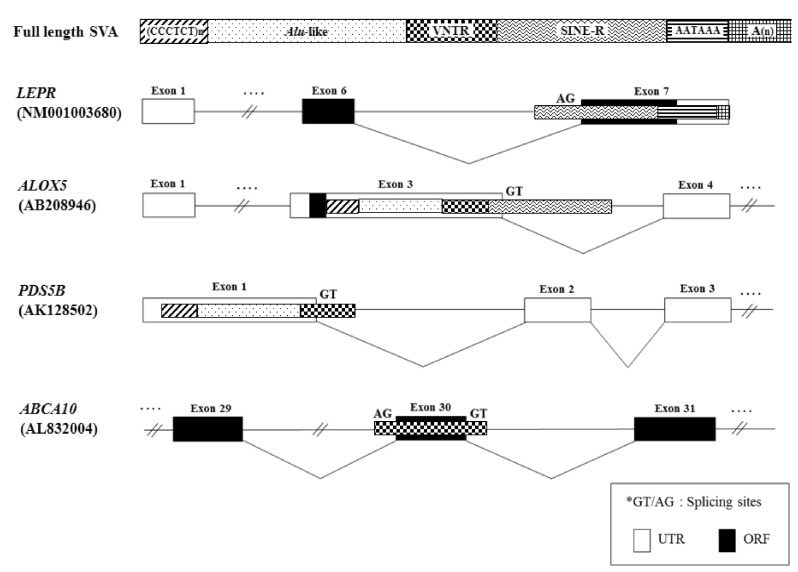
SVAs are composite elements consisting of multiple domains: a CCCTCT repeat, Alu-like domain, a GC-rich variable number of tandem repeat (VNTR), and SINE-R derived from the HERV-K LTR element [7-9, 21, 22]. They are flanked by target site duplications and terminate in a poly(A) tail (Fig. 1). In genomic sequence analysis, SVA elements are present in G + C-rich regions; however, they do not have any preferences for inter- or intragenic regions. SVA families are separated into six subfamilies (SVA-A to SVA-F), based upon point mutation and insertion and deletion events within the SINE-R [22]. Among them, four subfamilies (SVA-A, SVA-B, SVA-C, SVA-D) are present in gibbons and orangutans, while two subfamilies (SVA-E and SVA-F) are restricted to the human lineage [22].
SVA elements residing in genes are potentially disruptive in either orientation. Approximately 1/3 of all SVA elements in the human genome reside in genic regions, with 20% of those SVA elements being in the same orientation as a gene [23]. As shown in Fig. 2, we analyzed the genomic structure of SVA fusion genes using bioinformatic tools (Repeat-Masker program and University of California, Santa Cruz [UCSC] genome browser). SVA elements are detected in the 5'UTR of the HGSNAT (SVA-B, AK_057293), MRGPRX3 (SVA-D, NM_054031.2), HYAL1 (SVA-F, NM_153281), TCHH (SVA-F, AK_307946), and ATXN2L (SVA-F, AY_188334-8) genes, while some elements are observed in the 3'UTR of the SPICE1 (SVA-B, NM_144718), TDRKH (SVA-C, AK_225160), GOSR1 (SVA-D, NM_001007024), BBS5 (SVA-D, NM_1523842), NEK5 (SVA-D, NM_199289; AK_126330), ABHD2 (SVA-F, NM_007011), C1QTNF7 (SVA-F, NM_031911.3), ORC6L (SVA-F, AK_024077.1), TMEM69 (SVA-F, NM_016486.2), and CCDC137 (SVA-F, NM_199287.2) genes. Within the genic region, SVA elements seem to prefer the 3'UTR compared to another regions. Recently, a novel promoter derived from the SVA-D element was identified in the 5'UTR of the TBPL2 gene (DB089735) [19]. In order to understand whether the SVA element could act as a transcriptional regulator or not, the functional activity of the PARK7 SVA element was performed, suggesting that the SVA with the SINE-R region deleted showed significant activity to enhance reporter gene activity in SK-N-AS and MCF-7 cells [24]. Likewise, we have also demonstrated the activity of an alternative promoter by the HERV-H LTR element in the human GSDML gene [25]. The transcripts of the LTR-derived promoter were widely distributed in various tissues, whereas the transcripts of the original promoter were found in stomach tissues, compared to those of various human tissues [25]. A human-specific solitary LTR element (L47334) was previously shown to have enhancer activity in Tera-1 human testicular carcinoma cells [26]. The varying genetic structure of the alternative promoter or enhancer results in different functional effects [27]. Those events could regulate alternative transcript variants, resulting in the production of different protein isoforms [28]. In the case of leptin receptor isoform 219.1, SVA sequences form the C-terminal coding exon. Using the SVA-specific probe, Southern blot analysis in hominoids and Old World monkeys was performed, indicating that it was found to be a human-specific SVA element [29]. Furthermore, new exons are generated from intronic retrotransposons or the elements inserts into the exon, leading to a new transcript [30]. In the present study, the LEPR (SVA-C, NM_001003680), ALOX5 (SVA-D, AB_208946), PDS5B (SVA-D, AK_128502.2), and ABCA10 (SVA-F, AL_832004) genes showed alternative transcripts by the SVA exonization event. To investigate the supply of alternative splicing sites at SVA elements, we analyzed related splicing sites using human genomic sequences (http://genome.ucsc.edu/) (Fig. 3). In the case of ABCA10 (AL832004) genes, the SVA element provided both a splicing donor and acceptor site for the cassette exon. In the case of the ALOX5 (AB208946) and PDS5B (AK128502) genes, the SVA element provided a splicing donor site for alternative 5' splicing. Conversely, the LEPR (NM001003680) gene was provided a splicing acceptor site for alternative 3' splicing. These phenomena conformed to the canonical splicing site. Also, the PDS5B (AK128502) and ABCA10 (AL832004) genes induced exonization by providing alternative splicing sites in the VNTR region of SVA elements. Exonization of the LEPR (NM001003680) and ALOX5 (AB208946) genes occurred by the SINE-R region of SVA elements. This integration event of SVA elements into the exon or intron causes the promotion of transcriptional variants of functional genes. In WDR66 (NM_144668), the SVA-F element, inserted in intron 19, could produce human-specific transcripts that spliced to the last three exons [19]. Those transcripts, derived from SVA elements, could have important biological function in humans, therefore deserving further investigation in various tissues of hominoid primates.
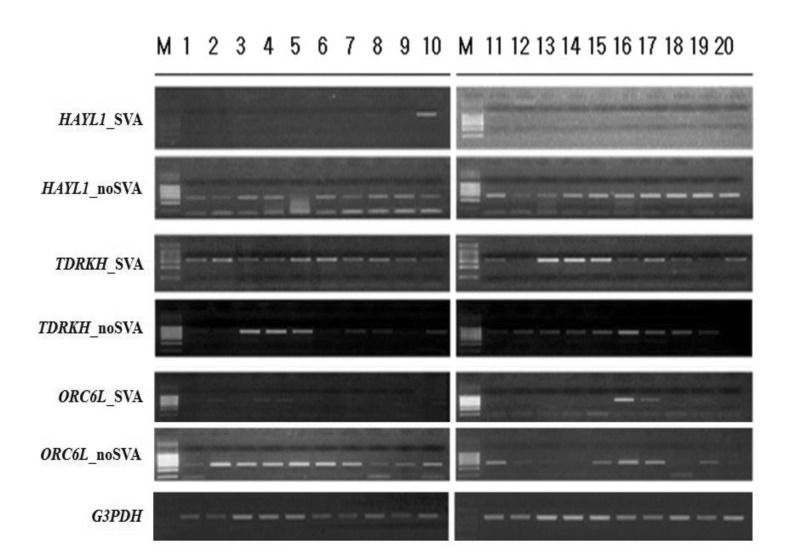
SVA elements are capable of generating individual variation in gene expression at loci in which they are present. RT-PCR amplification could be a good indicator to detect alternative variants derived from SVA elements. As shown in Fig. 4, dominant expression of HYAL1_SVA appeared in lung tissues, while HYAL1_noSVA showed ubiquitous expression in various human tissues. Expression of both transcripts (TDRKH_SVA and TDRKH_noSVA) of the TDRKH gene appeared to be ubiquitous. We also examined in silico expression of SVA fusion genes using microarray data obtained by the BioGPS database. Transcripts of the TCHH and HYAL1 genes appeared dominant in placenta and liver, respectively. The ALOX5 gene showed dominant expression in whole blood and lung. Also, dominant expression of the ABHD2 gene appeared in testis and prostate tissues, whereas the BBS5, NEK5, MRGPRX3, PDS5B, OR6W1P, ABCA10, TMEM69, C1QTNF7, and ATXN2L genes indicated ubiquitous expression (Fig. 5). SVA elements are nonautonomous retrotransposons that cause diseases in humans, and they are mobilized in trans by active L1 elements. The quantitative real-time polymerase chain reaction analysis of fukutin mRNA in lymphoblasts from FCMD patients indicated that the disease results from SVA-E insertion in the 3'UTR of the FUKUTIN gene. Sequence data demonstrated an abnormal splicing event by the integration of an SVA-E element [18]. In the case of the PMS2 gene, SVA-F has inserted in intron 7, and causes Lynch syndrome. Sequence analysis of the RT-PCR product revealed a 71-bp SVA-F element between PMS2 exon 7 and 8 in the aberrant transcripts [31]. Exon 3 of the PNPLA2 gene was interrupted by a 1.8-kb SVA-F insertion, which causes lipid storage disease with subclinical myopathy [32], while an SVA-F element inserted in intron 1 of the ARH gene causes hypercholesterolemia [33]. Taken together, alternative promoter, enhancer, polyadenylation, and exonization events by SVA elements cause various transcript isoforms and evolutionary dynamics that contribute to flexibility in the regulation of gene expression and hominoid radiation, including human disease.