 |
 |
- Search
| Genomics Inform > Volume 14(4); 2016 > Article |
Abstract
Environmental microbes like Bordetella petrii has been established as a causative agent for various infectious diseases in human. Again, development of drug resistance in B. petrii challenged to combat against the infection. Identification of potential drug target and proposing a novel lead compound against the pathogen has a great aid and value. In this study, bioinformatics tools and technology have been applied to suggest a potential drug target by screening the proteome information of B. petrii DSM 12804 (accession No. PRJNA28135) from genome database of National Centre for Biotechnology information. In this regards, the inhibitory effect of nine natural compounds like ajoene (Allium sativum), allicin (A. sativum), cinnamaldehyde (Cinnamomum cassia), curcumin (Curcuma longa), gallotannin (active component of green tea and red wine), isoorientin (Anthopterus wardii), isovitexin (A. wardii), neral (Melissa officinalis), and vitexin (A. wardii) have been acknowledged with anti-bacterial properties and hence tested against identified drug target of B. petrii by implicating computational approach. The in silico studies revealed the hypothesis that lpxD could be a potential drug target and with recommendation of a strong inhibitory effect of selected natural compounds against infection caused due to B. petrii, would be further validated through in vitro experiments.
Infectious diseases are major dreadful threat to human life, spread through many causative agents like bacteria, fungus, virus, etc. Therefore, development of potential drug is highly essential to fight against infectious diseases. Generally, methods of traditional drug design are tedious, time consuming, and cost intensive. Hence, several multidisciplinary approaches have gained research attention to reduce the time and cost during drug development process. In silico drug design is one of such approach, provides remarkable opportunity for identification of novel lead compounds as comparison to other conventional methods. Drug target identification is an important step involved in computer aided drug design process. In this context, the present study is planned to identify a potential drug target against infectious diseases caused by Bordetella petrii using various computational tools and techniques. B. petrii is a gram-negative coccobacilli of the phylum Proteobacteria, has been found exclusively in close association with humans and various warm-blooded animals due to its independent existence as an environmental facultative anaerobe [1]. B. petrii exhibits the host associated properties, and its strains have been detected from environmental samples like microbial consortia degrading aromatic compounds, marine sponges, polluted soil and a grass root consortium [2,3,4,5,6]. The genome of B. petrii contains 5,287,950 base pairs, and has been observed with greatest numbers of expressed virulence factors [7]. Infectious diseases like mandibular osteomyelitis [8], suppurative mastoiditis [9], chronic pulmonary diseases [10], respiratory infection in case of bronchiectasis and cystic fibrosis [11,12] have been reported due to B. petrii infection. Again, exposure to soil or water containing B. petrii may lead to local colonization and eventually serious nosocomial infections [13,14]. Some other bacteria like Escherichia coli [15], Pseudomonas aeruginosa [16], Chlamydophila pneumoniae [17], Staphylococcus aureus [18], and Haemophilus influenzae [19] have also been identified as a cause for such types of infections. Many Food and Drug Administration (FDA) approved drugs have been used against these pathogens, but those drugs have less effect on B. petrii due to development of substantial resistance [10], generated an opportunity to identify a potential drug target as well as to discover a novel lead compound against B. petrii using computational techniques. Again, due to adverse effect of most of the anti-infectious drugs of synthetic origin, natural compounds with anti-infectious properties have been tested in both in vivo and in vitro to be an alternative approach towards the treatment. In this context, some of the natural compounds such as ajoene (Allium sativum), allicin (A. sativum), cinnamaldehyde (Cinnamomum cassia), curcumin (Curcuma longa), gallotannin (active component of green tea and red wine), isoorientin (Anthopterus wardii), isovitexin (A. wardii), neral (Melissa officinalis), and vitexin (A. wardii) have been acknowledged with anti-bacterial properties [20,21,22,23,24,25]. Curcumin is a naturally occurring polyphenolic compound, a common component of Indian spice turmeric, having anti-tumor, anti-oxidant, and anti-inflammatory properties. It has been reported as an agent to reduce bacterial colony formation and inhibit lung cancer progression in high-risk chronic obstructive pulmonary disease (COPD) patients [20]. Similarly, gallotannin is an active component of green tea and red wine has been observed as a potent inhibitor of calcium-activated ClŌĆō channel, may be used for the treatment of cystic fibrosis [21]. Again, in vitro study of three flavone C-glycosides such as isoorientin, vitexin, and isovitexin isolated from A. wardii have been reported as potent inhibitors for interleukin-8, and matrix metalloproteinase-1, an inflammatory marker in case of COPD [22]. Moreover, most of the bacterial infection caused due to formation of bio film in the host. In this regard, an active compound cinnamaldehyde of C. cassia have been identified as a strong inhibitor against quorum sensing (QS) biofilm formation due to E. coli, and Vibrio harveyi [23] infection. Similarly, two extracts of A. sativum namely allicin and ajoene have been accounted for inhibitory effect against P. aeruginosa and E. coli [24]. Likewise, neral is an active component of M. officinalis, has been reported as anti-bacterial, anti-fungal, and anti-oxidant properties against P. aeruginosa, Klebsiella pneumoniae, S. aureus, and Citrobacter koseri infection [25], and may be applied for several infectious diseases including nosocomial infection. As, all of these nine natural compounds have been established with anti-bacterial properties, may be useful for treatment of B. petrii infection, encouraged authors to elucidate their inhibitory effect against the putative drug target of B. petrii through in silico approach.
The complete protein datasets of B. petrii (whole genome project accession No. PRJNA28135) were retrieved from Genome database of National Centre for Biotechnology Information (NCBI) web server (http://www.ncbi.nlm.nih. gov/genome/). The essentiality search [26] was carried out through the Database of Essential Gene (DEG) for screening out the essential proteins responsible for the survival of B. petrii in the host. The bit score parameter was considered as greater or equal to 100 for homology search using Basic Local Alignment Search Tool (BLAST) while accessing DEG (http://tubic.tju.edu.cn/deg/blast.php) database, to select bacterial essential proteins. The non-redundancy search was performed through CD-HIT (http://weizhong-lab.ucsd.edu/cdhit_suite/cgi-bin/index.cgi) web server to remove the redundant proteins among selected essential proteins of B. petrii. Further, by considering the resulted proteins of B. petrii, the sequence similarity search was carried out against human [27,28,29] followed by human gut flora microorganisms [30,31] using BLASTP algorithm to eliminate proteins homologous with human and human gut flora microorganisms respectively. During similarity search, the threshold for expected value (e-value) was fixed as greater or equal to 0.001. Furthermore, the protein sequences which are putative, unknown, hypothetical, probable, conserved hypothetical and unnamed protein were removed manually. The study of sub cellular localization was performed to identify the cytoplasm proteins among the selected proteins [32] using CELLO (http://cello.life.nctu.edu.tw/). Generally, proteins are annotated and ranked as per their evidence level in UniProt database which is also a factor to be considered while selecting putative drug targets. Afterwards, the Virulence Factor Database (VFDB) was explored, and the virulence proteins were screened out (http://www.mgc.ac.cn/VFs/blast/blast.html) among the selected targets of B. petrii through VFDB BLASTP search against VFDB core data sets with consideration of threshold e-value as greater than 0.0001 [31]. The above methodology is described through a flow diagram (Fig. 1A).
The participation of virulent proteins in any unique pathways of pathogen could be ensured and analyzed through the exploration of databases, is a common procedure to be followed while predicting putative drug targets. In this study, the pathway analysis was performed using Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.genome.jp/kegg/). Here, the exploration of KEGG database for pathway study was conducted twice. In the first case, the search was carried out for all obtained virulent proteins against existing pathways of B. petrii using KEGG BLAST algorithm. Secondly, unique pathways of B. petrii (pathways present in B. petrii, not in human) were sort listed from the KEGG database by comparing all pathways between B. petrii and human. Further, selection of common unique pathways [26,32,33] was made between the observed pathways of selected targets and obtained unique pathways of B. petrii. Furthermore, the selection pathways which are essential for the survival of the pathogen were made after detail inspection of obtained pathways. Finally, the key proteins regulating those pathways of B. petrii were considered as putative drug targets (Fig. 1B).
The putative drug targets of B. petrii obtained after pathway analysis were subjected for protein-protein network analysis using STRING (http://string-db.org/) web server. Here, the interaction between the drugs targets were quantified on the basis [34] of confidence value (0.007) and the level of interaction (maximum as 50). The drug targets observed as having higher numbers of interaction with other drug targets among the chosen data sets were considered for further analysis (Fig. 1B).
Similarly search was carried out for selected proteins using BLASTP algorithm against Protein Data Bank (PDB) database (http://www.rcsb.org/pdb/). The closest homolog for each selected target protein was considered as template based on lowest e-value and highest percentage of similarity The conserved domain region of potential drug target was predicted using conserved domain database (CDD) of NCBI web server (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) with default parameters (Fig. 1B).
Phyre2 (Protein Homology/Analogy Recognition Engine) server was used for modelling of three dimensional structure of putative drug target of B. petrii. The algorithm of Phyre2 server predict three dimensional structure of a protein sequence using the principles and techniques of homology and/or fold recognition, as structural folding pattern is more conserved in evolution than its amino acid sequence. Hence, a protein of interest (drug target) can be modelled with reasonable accuracy basing on a distantly related protein with known (template) structure [35]. The energetically stable structure was obtained through structural refinement and energy minimization using ModRefiner algorithm [36] of Zhang Lab web server (http://zhanglab.ccmb.med.umich.edu/ModRefiner/). The structural superimposition of refined model with template model was performed using Discovery Studio 3.5 suite. The structural quality and reliability of refined model were evaluated using RAMPAGE [37], ProSA-web [38], Protein Quality Predictor (ProQ) [39], and ERRAT [40] web tools.
The chemical structures of nine phytochemicals (ajoene, allicin, cinnamaldehyde, curcumin, gallotannin, isoorientin, isovitexin, neral, and vitexin) were downloaded from PubChem database of NCBI web server (http://ncbi.nlm.nih.gov/pccompound). The correction of structural geometry and energy minimization were carried out using Discovery Studio 3.5 suites.
Molecular docking between the predicted model of potential drug target with each of the nine natural compounds was conducted using AutoDock-4.2 algorithm (http://autodock.scripps.edu/), by considering the protein structure as rigid whereas ligands were kept as flexible. AutoDock algorithm requires pre calculated grid maps, one for each atom type present in the flexible molecules being docked, and it stores the potential energy arising from the interaction with rigid macro molecules [41]. Ligands were docked around the identified conserved region within the functional domain of the selected drug target. The favorable poses were selected on the basis of lowest binding energy calculated by AutoDock which indicates for most favorable binding conformation between receptor and ligands. Docking procedure was carried out as per normal methodology for protein-ligand docking [42,43]. The visualization and analysis of interaction between docked complexes were made using Discovery Studio 3.5 suites.
Availability of whole genome information of B. petrii DSM 12804 (accession No. PRJNA28135) in Genome database of NCBI web server encouraged authors to retrieve the entire set of 5,031 protein sequences in order to identify a potential drug target against it. Primarily, screening was made through identification of essential genes of B. petrii, considered as primitive strategy [26] for target identification. Here, out of total 5,031 proteins, only 1,841 were obtained as essential proteins and hence suspected as primary targets of B. petrii. To remove redundancy [29,30] among the putative drug targets, a non-redundancy search was carried out which resulted 1,802 proteins as non-redundant. Search for sequence homology with human proteins is an essential step to be followed for identification of proteins present only in pathogen but not in human [27,28,29]. Again, due to symbiotic relationship among human and human gut micro biota, homology search between proteins of pathogen and human gut flora microorganisms is also a crucial step [30,31] to be performed to eliminate the target proteins which are homologous to human gut flora microorganism. The resulted 799 numbers of proteins of B. petrii were not shown any significant similarity with human proteome as well as human gut flora proteome, hence were selected for downstream analysis. Furthermore, the reduction in protein dataset was done by selecting proteins reported as active and as well as functional. Out of 799 proteins, only 509 numbers of functionally characterized proteins were selected using the information available at UniProt database. As suggested by the literature, only cytoplasm proteins are suitable drug targets whereas membrane bound proteins are generally considered for vaccine development [32]. Among them only cytoplasm proteins (334 numbers) were considered for further study. Furthermore, screening was carried out on the basis of evidence report. As of UniProt database report, the evidence which supports the existence of proteins are classified as: (1) experimental evidence at protein level; (2) experimental evidence at transcript level; (3) protein inferred from homology; (4) protein predicted; and (5) protein uncertain. Out of these evidences, evidence at protein level is the best choice for protein existence, as it indicates the existence of the protein experimentally. But in the current investigation, not a single protein has been observed satisfying this criterion. Only 135 proteins which were inferred from homology subjected to virtual study. Identification of pathogenic proteins having virulence factor is also obligatory [31] while screening a drug target against any pathogenic microorganism. Therefore, homology search was carried out between 135 numbers of proteins and the core virulence proteins of B. petrii using VFDB BLAST. The result suggested, out of 135 selected proteins only 26 number of proteins found with virulence factor, which strongly supported for further study to predict potential drug target against B. petrii (Figs. 1 and 2).
The pathway analysis was conducted to screen out the putative drug targets and their involvement in important and unique pathways of the pathogen B. petrii which are essential for the survival of the pathogen. The preliminary search in KEGG database suggested for involvement of 22 proteins in different important pathways of B. petrii, and therefore the dataset of putative drug target reduced from 26 to 22. Further, exploration of all available pathways for both human and pathogen was conducted and compared, resulted 38 unique pathways present in B. petrii but absent in human. Furthermore, the comparison was made between the identified important pathways of B. petrii for 22 drug targets and 38 unique pathways identified in B. petrii, resulted 12 putative drug targets (Table 1) contributing towards the 3 common and unique pathways of B. petrii namely, lipopolysaccharide biosynthesis, two component system, and bacterial chemotaxis. As per literature suggest, lipopolysaccharide biosynthesis is an essential pathway [44] for the survival of B. petrii, a major component of the outer membrane of gram negative bacteria and referred as endotoxin. It is comprised of a hydrophilic polysaccharide and a hydrophobic component referred as lipid A which is responsible for major bio reactivity of endotoxin [44], also serves as a physical barrier providing protection towards bacteria from its surroundings [45]. Intense inspection on lipopolysaccharide biosynthesis pathway (Fig. 3) suggested, out of 12 selected putative drug targets (Table 1) only eight numbers (lpxH, kdsB, lpxK, lpxD, lpxB, kdsA, lpxC, and Bpet0453) were as key regulatory proteins [46]. Again, involvement of two components namely, amino acid and sugar metabolism and pentose phosphate pathway were found as regulating the complete pathway. lpxC, lpxD, lpxH, lpxB, and lpxK proteins are involved in amino acid and sugar metabolism, and among them lpxH was discarded as the synthesis of lpxB could alternatively possible by lpxD. Further, lpxB and lpxK were also rejected as potential drug target as the regulation of kdtA and waaA through lpxB and lpxK were otherwise possible by the proteins involved in pentose phosphate pathway. Furthermore, Bpet0453 (5.3.1.13), kdsA (2.5.1.55), and kdsB (2.7.7.38) proteins were found as key proteins of pentose phosphate metabolism components (Fig. 3), and regulating synthesis of kdtA and waaA, but the synthesis of same proteins was also control by proteins involved in amino acid and sugar metabolism to carried out the whole lipopolysaccharide biosynthesis. So the key proteins kdsA, kdsB, and Bpet0453 of pentose phosphate pathway were discarded. Finally, two proteins namely lpxD and lpxC were selected on the basis of their highly involvement in the regulation of lipopolysaccharide biosynthesis pathway than any other proteins (Figs. 2B and 3).
Protein network analysis is generally used for selection of drug targets having interaction with maximum numbers of other drug targets, an efficient technique to discover a potent drug target against pathogens [34]. Here, the biological interaction was studied for 2 obtained drug targets of B. petrii, and the assessment supported for both lpxD (with lpxC, lpxB, kdsA, kdsB, lpxH, lpxK, and bpet0453) and lpxC (with lpxD, lpxB, kdsA, kdsB, lpxK, and bpet0453) proteins showing interaction with other drug targets among the chosen data set (Table 2, Fig. 4). Therefore, lpxD and lpxC proteins were strongly recommended to be potential drug targets against the pathogen B. petrii.
Further, template structure identification for lpxD and lpxC proteins were undergone through similarity search against PDB. The template protein structure (PDB ID: 3PMO) from P. aeruginosa was selected for the drug target lpxD of B. petrii with 91% query coverage and 49% of identity where as template structure (PDB ID: 4FW3) from P. aeruginosa for lpxC protein of B. petrii with 88% query coverage and 56% of identity (Table 3). Though, it has been already established that lpxC catalyzes initially during the biosynthesis of the lipid A component of lipopolysaccharide and may be a potent drug target to block the lipopolysaccharide pathway of pathogens [47], the current investigation suggested, the status of lpxC protein B. petrii as un reviewed (Table 3) and have shown less number of interaction with other drug targets in comparison to lpxD (Table 2). Again, as per literature report, lpxD has a key role in biosynthesis of the lipid A component of lipopolysaccharide, and its inhibitors are also potentially attractive as antibacterial agents, as there is no close similarity found with human proteins [48]. Also, lpxD has a vital role in bio film formation which is causing nosocomial infection [16] and the status obtained from UniProt web server as reviewed (Table 3). This seems lpxD could be a potential drug target against B. petrii infection. Therefore, lpxD was proposed as a potent drug target subjected for further in silico study. Through BLAST search and inspection in CDD database, the conserved amino acid resides within the left-handed parallel beta helix (LbH) domain of lpxD were identified (Fig. 5A), containing three imperfect tandem repeats of a hexapeptide repeat motif (X-[STAV]-X-[LIV]-[GAED]-X).
Due to absence of experimental structure the tertiary model of lpxD (UniProt ID: A9INS9) protein was predicted by the single highest scoring template i.e., lpxD protein of P. aeruginosa (PDB ID: 3PMO, chain A) using phyre2 [35] web server with 100% confidence. Protein attains the lowest free energy during the native state; therefore the stability of the predicted model was improved through energy minimization [36] procedure, supported by the Root Mean Square Deviation score 0.612 obtained through the structural superimposition of lpxD models both before and after energy minimization (Fig. 5B). The refined lpxD protein model is publically available in Protein Model Database with PMDB ID (PM0080538).
The correctness and quality of predicted lpxD protein model of B. petrii was verified through RAMPAGE [37], ProSA-web [38], ProQ [39], and ERRAT [40] online server. The RAMPAGE analysis revealed 99.5% residues were folded properly suggesting good backbone folding pattern (Fig. 5C). Further, Z-score for lpxD model calculated by ProSA-web to be ŌĆō6.84 (Fig. 5D) which is well within the range of native conformation of experimental structures. Furthermore, ProQ computed a Levitt-Gerstein (LG) score of 2.785 and Max sub score to be 0.225 for the predicted model. A ProQ LG score >2.5 is necessary for suggesting a model is of good quality [39], implying high accuracy level of predicted structure. Also, the overall quality factor for lpxD model of B. petrii was predicted as 73.23% (Fig. 5E) by ERRAT web server. All of these evaluations recommended the reliability of the predicted lpxD model of B. petrii.
As per literature survey, nine natural compounds such as ajoene (A. sativum), allicin (A. sativum), cinnamaldehyde (C. cassia), curcumin (C. longa), gallotannin (active component of green tea and red wine), isoorientin (A. wardii), isovitexin(A. wardii), neral (M. officinalis), and vitexin (A. wardii) have been identified with medicinal properties and reported useful against different infectious diseases (Table 4, Fig. 6), and hence downloaded from PubChem database of NCBI web server.
The results of molecular docking were intended for energetically favorable binding poses for each nine natural compounds within the active pocket (LbH domain) of proposed drug target of B. petrii (Fig. 5A) as lowest binding energy and inhibition constant are generally preferred for favorable binding mode while docking small compounds with target protein molecule [42,43]. Again, among them isoorientin (ŌĆō5.66 kcal/mol), curcumin (ŌĆō5.38 kcal/mol), isovitexin (ŌĆō5.22 kcal/mol) found with lower binding energy and inhibition constants i.e., 71.35 ┬Ąm, 114.67 ┬Ąm, and 149.85 ┬Ąm, suggesting as strong inhibitors of lpxD protein. Further, the binding stability was proved through existence of hydrogen bonding pattern between the protein-ligand complexes (Table 5). Presence of hydrophobic amino acid residues within the binding site highly contributes towards the stability during molecular interaction between protein and small compound. Here, the resulted hydrophobic interaction between the docked complexes also supported for a strong binding affinity of natural compounds towards the proposed drug target (lpxD) of B. petrii (Table 6, Fig. 7). Also, the presence of some positively and negatively charged amino acid residues of lpxD within 4 ├ģ distance from the ligands suggested for electrostatic force of interaction between docked complexes (Table 6, Fig. 7). Again, among nine active natural compounds, isoorientin (ŌĆō5.66 kcal/mol) was observed as the most suitable inhibitor with lowest binding energy and inhibition constant (71.35 ┬Ąm), and formed hydrogen bond with amino acid residues like Ala 186, Glu 149, and Gly 227 of lpxD (Table 5, Fig. 7F). Strong hydrophobic interactions were seen due to presence of Val 165, Ala 186, and Leu 226 within 4 ├ģ distance from the ligand (Table 6, Fig. 7F). All of these above observation established isoorientin to be a strong inhibitor against lpxD of B. petrii.
In conclusions, Identification of potential drug target has a vital role in designing a novel drug against any infectious disease. B. petrii strains have been reported as causing infections in humans and various warm blooded animals. In the present investigation, authors attempted to narrow down the complete data set of known 5,031 protein sequences of B. petrii DSM 12804 (accession No. PRJNA28135) in order to identify a potential drug target computationally. In silico study revealed lpxD protein of B. petrii as a potential drug target, is an essential protein of the pathogen and involved in the biosynthesis of the lipid A component of lipopolysaccharide. Again, the observations made through computational study also strongly supported lpxD to be a potential drug target due to the absence of lpxD protein in any human essential pathways as well as without any significant percentage of sequence homology with human and human gut flora micro biota. Again, previous literature reports also established the involvement of lpxD in nosocomial infections caused due to pathogens and hence, could be used as a potential drug target [15,16] against B. petrii. Furthermore, due to adverse effect of synthetic drugs, the uses of active natural compounds have gained an increasing attention towards the treatment or prevention of infectious diseases. Moreover, some of the phytochemicals like ajoene (garlic), allicin (ginger), cinnamaldehyde (Chinese cinnamon), curcumin (turmeric), gallotannin (green tea and red wine), isoorientin (blueberry relative found in Central and South America), isovitexin (blueberry relative found in Central and South America), neral (lemon balm), and vitexin (blueberry relative found in Central and South America) have been reported as having anti-oxidant, anti-inflammatory, and anti-bacterial properties, might be used against COPD, cystic fibrosis, QS biofilm formation, nosocomial infection, and other types of bacterial infection [20,21,22,23,24,25]. Therefore, authors excited to apply molecular docking approach to check the inhibitory effects of above mentioned nine natural compounds (Table 4, Fig. 6) against the proposed drug target (lpxD) of B. petrii. The docking study results also supported for the confirmation of the mechanism of blocking of lipid A biosynthesis in B. petrii which is essential for the survival of the bacterium. Again, isoorientin was noticed with highest inhibitory effect among nine natural compounds supported by lowest binding energy and inhibition constant (Table 5). Binding stability was also confirmed through observation of strong hydrophobic, electrostatic and hydrogen bonding interactions between protein-ligand complexes (Tables 5 and 6, Fig. 7F). Hence, isoorientin could be suggested as most suitable natural compounds against the proposed drug target (lpxD) of B. petrii. The overall hypothesis generated through implementation of in silico approach would be a great aid and value towards designing a novel drug against B. petrii infection and which would be further validated through in vitro experiments.
Acknowledgments
The authors acknowledge Department of Biotechnology, Ministry of Science and Technology, Government of India for providing necessary infrastructure facility to carry out this research.
References
1. von Wintzingerode F, Schattke A, Siddiqui RA, R├Čsick U, G├Čbel UB, Gross R. Bordetella petrii sp. nov., isolated from an anaerobic bioreactor, and emended description of the genus Bordetella. Int J Syst Evol Microbiol 2001;51(Pt 4):1257ŌĆō1265. PMID: 11491321.


2. Wang F, Grundmann S, Schmid M, D├Črfler U, Roherer S, Charles Munch J, et al. Isolation and characterization of 1,2,4-trichlorobenzene mineralizing Bordetella sp. and its bioremediation potential in soil. Chemosphere 2007;67:896ŌĆō902. PMID: 17204305.


3. Bianchi F, Careri M, Mustat L, Malcevschi A, Musci M. Bioremediation of toluene and naphthalene: development and validation of a GC-FID method for their monitoring. Ann Chim 2005;95:515ŌĆō524. PMID: 16235785.


4. Sfanos K, Harmody D, Dang P, Ledger A, Pomponi S, McCarthy P, et al. A molecular systematic survey of cultured microbial associates of deep-water marine invertebrates. Syst Appl Microbiol 2005;28:242ŌĆō264. PMID: 15900971.


5. Chowdhury SP, Schmid M, Hartmann A, Tripathi AK. Identification of diazotrophs in the culturable bacterial community associated with roots of Lasiurus sindicus, a perennial grass of Thar Desert, India. Microb Ecol 2007;54:82ŌĆō90. PMID: 17264993.


6. Gross R, Guzman CA, Sebaihia M, dos Santos VA, Pieper DH, Koebnik R, et al. The missing link: Bordetella petrii is endowed with both the metabolic versatility of environmental bacteria and virulence traits of pathogenic Bordetellae. BMC Genomics 2008;9:449. PMID: 18826580.



7. Nogi M, Bankowski MJ, Pien FD. Septic arthritis and osteomyelitis due to Bordetella petrii. J Clin Microbiol 2015;53:1024ŌĆō1027. PMID: 25540393.



8. Fry NK, Duncan J, Malnick H, Warner M, Smith AJ, Jackson MS, et al. Bordetella petrii clinical isolate. Emerg Infect Dis 2005;11:1131ŌĆō1133. PMID: 16022798.



9. Stark D, Riley LA, Harkness J, Marriott D. Bordetella petrii from a clinical sample in Australia: isolation and molecular identification. J Med Microbiol 2007;56(Pt 3):435ŌĆō437. PMID: 17314377.


10. Le Coustumier A, Njamkepo E, Cattoir V, Guillot S, Guiso N. Bordetella petrii infection with long-lasting persistence in human. Emerg Infect Dis 2011;17:612ŌĆō618. PMID: 21470449.



11. Spilker T, Liwienski AA, LiPuma JJ. Identification of Bordetella spp. in respiratory specimens from individuals with cystic fibrosis. Clin Microbiol Infect 2008;14:504ŌĆō506. PMID: 18325036.


12. Moissenet D, Bingen E, Arlet G, Vu-Thien H. Use of 16S rRNA gene sequencing for identification of ŌĆ£Pseudomonas-likeŌĆØ isolates from sputum of patients with cystic fibrosis. Pathol Biol (Paris) 2005;53:500ŌĆō502. PMID: 16081224.


13. G├│mez-Cerezo J, Su├Īrez I, R├Łos JJ, Pe├▒a P, Garc├Ła de Miguel MJ, de Jos├® M, et al. Achromobacter xylosoxidans bacteremia: a 10-year analysis of 54 cases. Eur J Clin Microbiol Infect Dis 2003;22:360ŌĆō363. PMID: 12750959.


14. Conway SP, Brownlee KG, Denton M, Peckham DG. Antibiotic treatment of multidrug-resistant organisms in cystic fibrosis. Am J Respir Med 2003;2:321ŌĆō332. PMID: 14719998.


15. Buetow L, Smith TK, Dawson A, Fyffe S, Hunter WN. Structure and reactivity of LpxD, the N-acyltransferase of lipid A biosynthesis. Proc Natl Acad Sci U S A 2007;104:4321ŌĆō4326. PMID: 17360522.



16. Alshalchi SA, Anderson GG. Expression of the lipopolysaccharide biosynthesis gene lpxD affects biofilm formation of Pseudomonas aeruginosa. Arch Microbiol 2015;197:135ŌĆō145. PMID: 25173672.


17. Ravindranath BS, Krishnamurthy V, Krishna V, Vasudevanayaka KB. In silico analyses of metabolic pathway and protein interaction network for identification of next gen therapeutic targets in Chlamydophila pneumoniae. Bioinformation 2013;9:605ŌĆō609. PMID: 23904736.



18. LeFrock J, Mader J, Smith B, Carr B. Bone and joint infections caused by gram-positive bacteria: treatment with cefotaxime. Infection 1985;13(Suppl 1):S50ŌĆōS55. PMID: 4055056.


19. Hu J, Wang X, Ai T, Xie X, Liu X, Liu H, et al. Multicenter prospective epidemiological studies on Haemophilus influenzae infection among hospitalized children with lower respiratory tract infections. Zhonghua Er Ke Za Zhi 2016;54:119ŌĆō125. PMID: 26875462.

20. Moghaddam SJ, Barta P, Mirabolfathinejad SG, Ammar-Aouchiche Z, Garza NT, Vo TT, et al. Curcumin inhibits COPD-like airway inflammation and lung cancer progression in mice. Carcinogenesis 2009;30:1949ŌĆō1956. PMID: 19793800.



21. Namkung W, Thiagarajah JR, Phuan PW, Verkman AS. Inhibition of Ca2+-activated Cl-channels by gallotannins as a possible molecular basis for health benefits of red wine and green tea. FASEB J 2010;24:4178ŌĆō4186. PMID: 20581223.



22. Flores G, Dastmalchi K, Dabo AJ, Whalen K, Pedraza-Pe├▒alosa P, Foronjy RF, et al. Antioxidants of therapeutic relevance in COPD from the neotropical blueberry Anthopterus wardii. Food Chem 2012;131:119ŌĆō125. PMID: 22363097.



23. Niu C, Afre S, Gilbert ES. Subinhibitory concentrations of cinnamaldehyde interfere with quorum sensing. Lett Appl Microbiol 2006;43:489ŌĆō494. PMID: 17032221.


24. Niu C, Gilbert ES. Colorimetric method for identifying plant essential oil components that affect biofilm formation and structure. Appl Environ Microbiol 2004;70:6951ŌĆō6956. PMID: 15574886.



25. Jalal Z, El Atki Y, Lyoussi B, Abdellaoui A. Phytochemistry of the essential oil of Melissa officinalis L. growing wild in Morocco: preventive approach against nosocomial infections. Asian Pac J Trop Biomed 2015;5:458ŌĆō461.

26. Damte D, Suh JW, Lee SJ, Yohannes SB, Hossain MA, Park SC. Putative drug and vaccine target protein identification using comparative genomic analysis of KEGG annotated metabolic pathways of Mycoplasma hyopneumoniae. Genomics 2013;102:47ŌĆō56. PMID: 23628646.


27. Haag NL, Velk KK, Wu C. Potential antibacterial targets in bacterial central metabolism. Int J Adv Life Sci 2012;4:21ŌĆō32. PMID: 24151543.


28. Sakharkar KR, Sakharkar MK, Chow VT. A novel genomics approach for the identification of drug targets in pathogens, with special reference to Pseudomonas aeruginosa. In Silico Biol 2004;4:355ŌĆō360. PMID: 15724285.

29. Vetrivel U, Subramanian G, Dorairaj S. A novel in silico approach to identify potential therapeutic targets in human bacterial pathogens. Hugo J 2011;5:25ŌĆō34. PMID: 23205162.




30. Pandey M, Tiwari A, Maurya S, Singh D, Srivastava P. Choke point analysis with subtractive proteomic approach for insilico identification of potential drug targets in Shigella dysenteriae. Int J Comput Appl 2015;109:29ŌĆō34.

31. Shanmugham B, Pan A. Identification and characterization of potential therapeutic candidates in emerging human pathogen Mycobacterium abscessus: a novel hierarchical in silico approach. PLoS One 2013;8:e59126. PMID: 23527108.



32. Barh D, Kumar A. In silico identification of candidate drug and vaccine targets from various pathways in Neisseria gonorrhoeae. In Silico Biol 2009;9:225ŌĆō231. PMID: 20109152.


33. Anishetty S, Pulimi M, Pennathur G. Potential drug targets in Mycobacterium tuberculosis through metabolic pathway analysis. Comput Biol Chem 2005;29:368ŌĆō378. PMID: 16213791.


34. Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 2011;39:D561ŌĆōD568. PMID: 21045058.



35. Kelley LA, Sternberg MJ. Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 2009;4:363ŌĆō371. PMID: 19247286.


36. Xu D, Zhang Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys J 2011;101:2525ŌĆō2534. PMID: 22098752.



37. Lovell SC, Davis IW, Arendall WB 3rd, de Bakker PI, Word JM, Prisant MG, et al. Structure validation by Calpha geometry: phi,psi and Cbeta deviation. Proteins 2003;50:437ŌĆō450. PMID: 12557186.


38. Wiederstein M, Sippl MJ. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 2007;35:W407ŌĆōW410. PMID: 17517781.



39. Wallner B, Elofsson A. Can correct protein models be identified? Protein Sci 2003;12:1073ŌĆō1086. PMID: 12717029.



40. Colovos C, Yeates TO. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 1993;2:1511ŌĆō1519. PMID: 8401235.



41. Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 2009;30:2785ŌĆō2791. PMID: 19399780.



42. Kumar S, Jena L, Galande S, Daf S, Mohod K, Varma AK. Elucidating molecular interactions of natural inhibitors with HPV-16 E6 oncoprotein through docking analysis. Genomics Inform 2014;12:64ŌĆō70. PMID: 25031569.



43. Jagadeb M, Konkimalla VB, Rath SN, Das RP. Elucidation of the inhibitory effect of phytochemicals with Kir6.2 wild-type and mutant models associated in type-1 diabetes through molecular docking approach. Genomics Inform 2014;12:283ŌĆō288. PMID: 25705171.



44. Wang X, Quinn PJ. Lipopolysaccharide: biosynthetic pathway and structure modification. Prog Lipid Res 2010;49:97ŌĆō107. PMID: 19815028.


45. Rosenfeld Y, Shai Y. Lipopolysaccharide (Endotoxin)-host defense antibacterial peptides interactions: role in bacterial resistance and prevention of sepsis. Biochim Biophys Acta 2006;1758:1513ŌĆō1522. PMID: 16854372.


46. Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 2016;44(D1):D457ŌĆōD462. PMID: 26476454.



47. Barb AW, Zhou P. Mechanism and inhibition of LpxC: an essential zinc-dependent deacetylase of bacterial lipid A synthesis. Curr Pharm Biotechnol 2008;9:9ŌĆō15. PMID: 18289052.



Fig.┬Ā1
The methodology of identification of potential drug target of Bordetella petrii described through flow diagram. The methodology of subtractive proteomics deciphered in panel A, and rest in panel B.
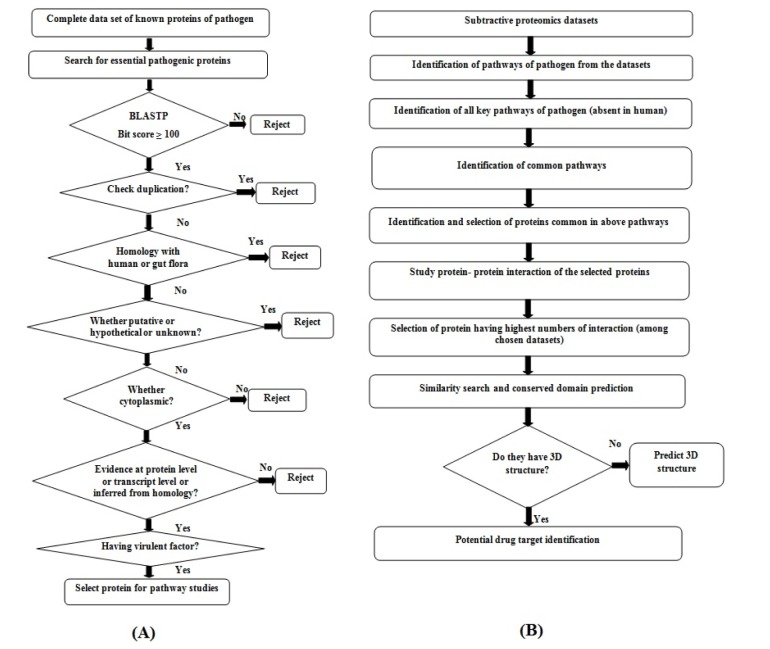
Fig.┬Ā2
The results of screening putative drug targets in Bordetella petrii DSM 12804 deciphered (A), where numbers in left hand side indicates the identified drug targets after completion of each step. (B) The resulted drug targets in each step represented using bar chart. DEG, Database of Essential Gene; BLAST, Basic Local Alignment Search Tool; VFDB, Virulence Factor Database; KEGG, Kyoto Encyclopedia of Genes and Genomes.

Fig.┬Ā3
The lipopolysaccharide biosynthesis pathway of Bordetella petrii explored from Kyoto Encyclopedia of Genes and Genomes (KEGG) [46] web server. The key protein lpxD (KEGG ID: Bpet2525) in the pathway highlighted in red color.

Fig.┬Ā4
The protein-protein network study of drug targets namely, lpxD, and lpxC depicted in section (A) and (B) respectively.

Fig.┬Ā5
The conserved amino acid residues within left-handed parallel beta helix (LbH) domain of lpxD protein between Bordetella petrii and Pseudomonas aeruginosa (PDB ID: 3PMO, Chain A) are identified and deciphered (A). The structural super imposition between identified drug target lpxD (red color) of B. petrii and template (PDB ID: 3pmo, Chain A) (blue color) of P. aeruginosa represented (B). The backbone folding pattern was studied through phi and psi torsion angle in Ramachandran plot represented (C). The overall model quality evaluated and deciphered (D), where the black dot in the plot represents for predicted structural model of lpxD, falls within the range of X-ray and NMR model quality. The errors within the non-bonded interaction plotted (E), the amino acid residues and percentage of error represented in X-axis and Y-axis respectively, the deep black color represents error regions in corresponding amino acid residues during non-bonded interaction.
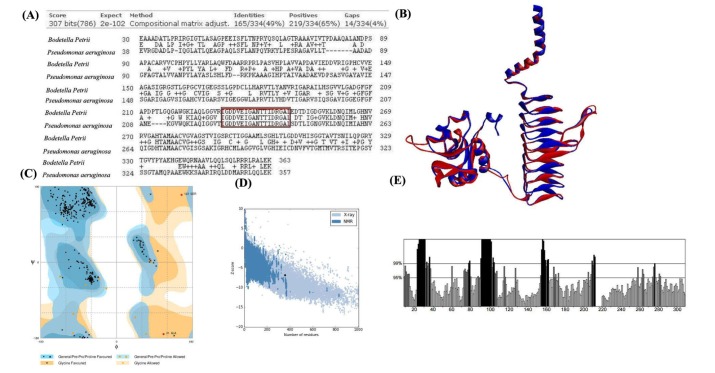
Fig.┬Ā6
Chemical structures of nine natural compounds: ajoene (A), allicin (B), cinnamaldehyde (C), curcumin (D), gallotannin (E), isoorientin (F), isovitexin (G), neral (H), and vitexin (I).
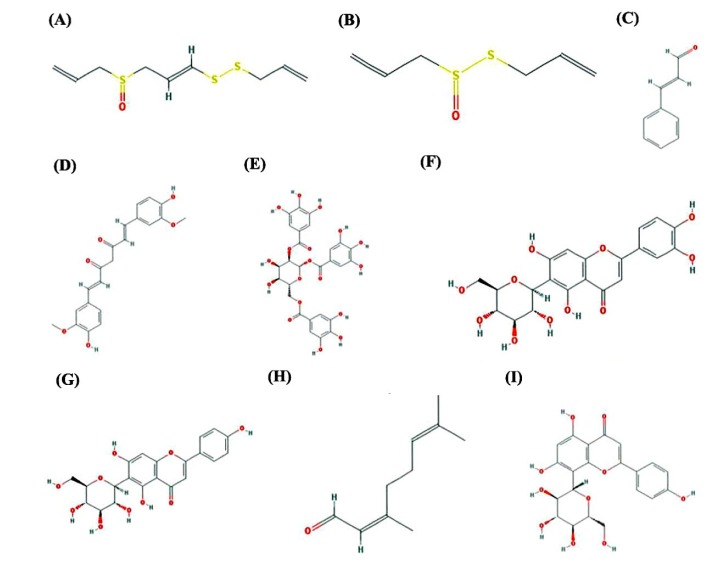
Fig.┬Ā7
Interaction profile of nine natural compounds: ajoene (A), allicin (B), cinnamaldehyde (C), curcumin (D), gallotanin (E), isoorientin (F), isovitexin (G), neral (H), and vitexin (I) with proposed drug target (lpxD protein) of Bordetella petrii. Amino acid residues participated in hydrogen bonding and interacting within a distance of 4├ģ in the active pocket of LbH domain of lpxD protein with natural compounds was deciphered here.

Table┬Ā1.
Twelve putative and virulent drug targets associated in three unique pathways of Bordetella petrii
Table┬Ā2.
Protein-protein interaction of selected drug targets of Bordetella petrii
Table┬Ā3.
Two proposed drug targets (lpxD, lpxC) of Bordetella petrii along with their homologous
Table┬Ā4.
Collected information about selected nine natural compounds against Bordetella petrii
| Serial No. | Natural compounds | PubChem compound ID | Molecular weight (g/mol) | Natural source | Disease | Reference |
|---|---|---|---|---|---|---|
| 1 | Ajoene | 5386591 | 234.40186 | Garlic | Quorum sensing inhibition of Pseudomonas aeruginosa and Escherichia coli | [24] |
| 2 | Allicin | 65036 | 162.273 | Ginger | Quorum sensing inhibition of P. aeruginosa and E. coli | [24] |
| 3 | Cinnamaldehyde | 637511 | 132.15922 | Chinese cinnamon | Biofilm formation | [23] |
| 4 | Curcumin | 969516 | 368.3799 | Turmeric | Pulmonary disease | [20] |
| 5 | Gallotannin | 12795683 | 636.46866 | Green tea and red wine | Cystic fibrosis | [21] |
| 6 | Isoorientin | 114776 | 448.3769 | Blueberry relative found in Central and South America | Chronic obstructive pulmonary disease | [22] |
| 7 | Isovitexin | 162350 | 432.3775 | Blueberry relative found in Central and South America | Chronic obstructive pulmonary disease | [22] |
| 8 | Neral | 643779 | 152.23344 | Lemon balm | Nosocomial infection | [25] |
| 9 | Vitexin | 5280441 | 432.3775 | Blueberry relative found in Central and South America | Chronic obstructive pulmonary disease | [22] |
Table┬Ā5.
Polar contact information obtained from docking calculation between selected natural compounds and drug target (lpxD) of Bordetella petrii
Table┬Ā6.
Close amino acid residues of drug target (lpxD) of Bordetella petrii participating during interaction with nine natural compounds within a distance of 4 ├ģ
| Serial No. | Natural compounds | Amino acids within 4 ├ģ |
Amino acid residues |
|||
|---|---|---|---|---|---|---|
| Hydrophobic interactionsa | Hydrogen bondingb | Positively chargedc | Negatively chargedd | |||
| 1 | Ajoene | ARG 189, GLY 191, ALA 192, ARG 230, ILE 231, GLY 232, ASP 233, ASP 252, ILE 253, GLY 254 | ALA 192, ILE 231, ILE 253 | ASP 233 | ARG 189, ARG 230 | ASP 233, ASP 252 |
| 2 | Allicin | PRO 118, ARG 119, LEU 120, ASP 139, VAL 140, ARG 141, GLY 157, SER 158, THR 159, ASP 175, CYS 176, LEU 177 | LEU 120, VAL 1 40, LEU 177 | PRO 118 | ARG 119, ARG 141 | ASP 139, ASP 175 |
| 3 | Cinnamaldehyde | ASN 187, ASP 205, ALA 224, GLN 225, GLY 227, ARG 245 | ALA 224 | GLN 225, ARG 245 | ARG 245 | ASP 205 |
| 4 | Curcumin | ARG 230, ILE 231, GLY 232, ASP 233, ASP 252, ILE 253, GLY 254, ASP 255, ARG 270, VAL 271, GLY 272, VAL 288, ILE 289, GLY 290, SER 291, LEU 307, ASP 309 | ILE 231, ILE 253, VAL 271, VAL 288, ILE 289, LEU 307 | VAL 271, ASP 309 | ARG 230, ARG 270 | ASP 233, ASP 252, ASP 309 |
| 5 | Gallotannin | GLU 149, VAL 165, ILE 166, GLY 167, GLU 168, GLY 169, LEU 184, TYR 185, ALA 186, ASN 187, GLY 203, GLN 225, LEU 226, GLY 227, GLY 228, ARG 245 | VAL 165, ILE 166, LEU 184, ALA 186, LEU 226 | ALA 186, AGR 245, LEU 184, ASN 187, GLY 227 | ARG 245 | GLU 149, GLU 168 |
| 6 | Isoorientin | GLU 149, VAL 165, GLY 167, GLU 168, THR 183, TYR 185, ALA 186, ASN 187, LEU 226, GLY 227, GLY 228, ARG 245 | VAL 165, ALA 186, LEU 226 | ALA 186, GLU 149, GLY 227 | ARG 245 | GLU 149, GLU 168 |
| 7 | Isovitexin | GLU 340, ARG 343, ASN 344, VAL 347, LEU 348, GLN 349, GLN 350, LEU 351, SER 352, ARG 355 | VAL 347, LEU 348, LEU 351 | ASN 344, SER 352, ARG 355, LEU 348 | ARG 343, ARG 355 | GLU 340 |
| 8 | Neral | ARG 189, GLY 191, ALA 192, ARG 230, ILE 231, GLY 232, ASP 233, ASP 252, ILE 253, GLY 254 | ALA 192, ILE 231 | ASP 233 | ARG 189, ARG 230 | ASP 233, ASP 252 |
| 9 | Vitexin | SER 171, LEU 172, GLY 173, PRO 174 ARG 189, ILE 190, GLY 191, ALA 192, ARG 230, ILE 231, GLY 232, ASP 233 | LEU 172 | SER 171, ARG 189 | ARG 189, ARG 230 | ASP 233 |
- TOOLS
-
METRICS

-
- 8 Crossref
- 0 Scopus
- 9,696 View
- 124 Download





