 |
 |
- Search
| Genomics Inform > Volume 17(1); 2019 > Article |
|
Abstract
The chinstrap (Pygoscelis antarcticus) and gentoo (P. papua) penguins are distributed throughout Antarctica and the sub-Antarctic islands. In this study, high-quality de novo assemblies of blood transcriptomes from these penguins were generated using the Illumina MiSeq platform. A total of 22.2 and 21.8 raw reads were obtained from chinstrap and gentoo penguins, respectively. These reads were assembled using the Oases assembly platform and resulted in 26,036 and 21,854 contigs with N50 values of 929 and 933 base pairs, respectively. Functional gene annotations through pathway analyses of the Gene Ontology, EuKaryotic Orthologous Groups, and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases were performed for each blood transcriptome, resulting in a similar compositional order between the two transcriptomes. Ortholog comparisons with previously published transcriptomes from the Adélie (P. adeliae) and emperor (Aptenodytes forsteri) penguins revealed that a high proportion of the four penguins’ transcriptomes had significant sequence homology. Because blood and tissues of penguins have been used to monitor pollution in Antarctica, immune parameters in blood could be important indicators for understanding the health status of penguins and other Antarctic animals. In the blood transcriptomes, KEGG analyses detected many essential genes involved in the major innate immunity pathways, which are key metabolic pathways for maintaining homeostasis against exogenous infections or toxins. Blood transcriptome studies such as this may be useful for checking the immune and health status of penguins without sacrifice.
Penguins are a monophyletic group within the Spheniscidae family [1]. Approximately 18 species breed sympatrically in several regions within Antarctica and the sub-Antarctic islands [2]. Pygoscelids include three species of penguin (Adélie Pygoscelis adeliae, chinstrap P. antarctica, and gentoo P. papua), and represent congeneric penguins based on their co-occurrence, coexistence, and similarities in their breeding and foraging ecologies [3-5]. Pygoscelids are densely distributed throughout the South Shetland and South Orkney Islands between the latitudes 54°S and 65°S. The chinstraps have a more southern distribution and are located almost exclusively around the Antarctic Peninsula [1,6]. The gentoo have the most northern distribution and forage deeper in the water column. The Adélie and chinstrap are relatively ice-tolerant and depend on ice more than the gentoo [1,6]. In 2017, pygoscelids were placed on the International Union for Conservation of Nature Red List of Threatened Species at the level of Least Concern (http://www.iucnredlist.org). Because their habitats are strongly correlated with Antarctic environmental conditions, fluctuations in the marine environment due to global warming could be the most obvious factor influencing their declining populations [7]. Therefore, consistent monitoring and conservation of pygoscelids is highly recommended.
Penguins serve as important models for studying ecology, the environment, physiology, behavior, reproduction, and population dynamics [8,9]. Antarctica is continuously threated by anthropogenic activities affecting the atmosphere and water including climate change, over-fishing, disease, and tourism [10-12]. Penguins are promising sentinels for monitoring marine pollution and disease in Antarctica given their high trophic position, long lifespan, philopatry, and conspicuous nature [8,9]. While previous studies have analyzed pathogen diversity or toxicant accumulation from penguin blood samples [13-18], to date there have been no published gene expression studies from the blood of any penguin species.
In this study, we analyzed the blood transcriptome of chinstrap (P. antarcticus) and gentoo (P. papua) penguins using Illumina MiSeq sequencing and a series of bioinformatics tools. Given that blood is highly sensitive to diverse endogenous and exogenous stimulations, including parasites, pathogens, and environmental pollutants, transcriptomic profiling and the measurement of target gene/pathway-specific expression is useful for predicting homeostasis, disease, immunity, and population health status in penguins. Although genomic analyses of penguin blood have been limited, several analytical applications have been conducted on their blood tissues. For example, blood-borne parasitic infections are a potential factor for the mortality of wild penguins [19-21]. Chemical analyses (i.e., dioxins, persistent organic pollutants, polychlorinated biphenyls, and pesticides) of blood samples from penguins have been used as a promising indicator of Antarctic environmental contamination [22-25]. In fact, among birds, blood transcriptome profiling has been successfully employed to predict the homeostasis of immune systems and to analyze transcriptional changes after pathogen or pollutant challenges [26-30]. Thus, genomic information from penguin blood samples will allow the evaluation of their health status using gene expression analyses to determine their immune status.
All animal handling and experimental procedures were approved by the Animal Welfare Ethical Committee and the Animal Experimental Ethics Committee of the Korea Polar Research Institute (KOPRI).
Blood specimens (approximately 1 mL) were collected from one individual of each penguin species on King George Island, South Shetland Islands, Antarctica (Table 1). Both species were identified in reference to morphological characteristics and their mitochondrial cytochrome oxidase subunit 1 sequences. Blood samples were immediately transferred to a vial containing RNAlater (Qiagen, Valencia, CA, USA) and stored at –20°C until RNA extraction. Total RNA was extracted using an RNeasy Micro Kit (Qiagen) according to the manufacturer’s instructions and stored in RNAstable (Biometrica, San Diego, CA, USA). The quality and quantity of the total RNA was assessed using the Bioanalyer 2100 (Agilent Technologies, Santa Clara, CA, USA). High-quality mRNA (2 µg) was used to generate a double-stranded cDNA library using poly A selection. Entire experimental reagents and equipments were purchased from Illumina Inc. (San Diego, CA, USA). The NuGEN Encore Complete RNA-Seq Library System (NuGEN, San Carlos, CA, USA) was used to construct paired-end libraries (PE500) of sheared cDNA (500 bp fragments) that were sequenced on an Illumina MiSeq System platform (300 × 2 paired-end reads). Index and adaptor sequences were trimmed using Trimmomatic [31] and low-quality reads were removed using the FASTX toolkit [32] with parameters set to-t = 20,-l = 70, and-Q = 33.
Low-quality reads (average quality score < 10), adapters, linkers, and polymerase chain reaction primers were removed through quality filtering. High-quality reads were assembled de novo using Oases (ver. 0.2.08) with default parameters [33]. The TransDecoder was used to identify coding regions. Unclustered transcripts and the longest sequences were considered unigenes. To identify functional transcripts, all unigenes were searched against the National Center for Biotechnology Information (NCBI) nonredundant database using BLASTx with an E-value threshold of 1.00E-04. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses of all contigs were performed using the Blast2GO sequence annotation tool (ver. 4.0) [34]. BLAST searches and functional domain annotations were conducted using InterProScan within the Blast2GO software package.
Finally, the assembled data were arranged in terms of read length, gene annotation, GenBank number, E-value, species, and species accession number. The mRNA expression levels were calculated using the reads per kilobase of the transcriptome per million mapped reads (RPKM) method [35].
Raw RNA-seq data were deposited into the NCBI Sequence Read Archive (accession numbers, SRR7874621 for P. antarcticus and SRR7874380 for P. papua) under the bioproject numbers, PRJNA491756 for P. antarcticus and PRJNA491755 for P. papua, respectively. The Transcriptome Shotgun Assembly project was deposited into the DDBJ/ENA/GenBank database under accession numbers, GGXL00000000 for P. antarcticus and GGXK00000000 for P. papua.
Approximately 22.2 and 21.8 million reads were obtained by Illumina sequencing for P. antarcticus (accumulated base pairs, 6,675,343,822 bp) and P. papua (accumulated base pairs, 6,574,802,598 bp), respectively. In total, 26,036 contigs with an average length of 863 bp and an N50 length of 929 bp were obtained in the P. antarcticus blood transcriptome, whereas in the P. papua blood transcriptome, 21,854 contigs with an average length of 857 bp and an N50 length of 933 bp were obtained.
Overall, the principal BLAST hits of the penguin transcripts exhibited high similarity to avian genomic information at both class and family levels (Fig. 1A). Of the top hits, 86% of P. antarcticus and P. papua contigs were homologous to transcripts from the class Aves. At the family level, both contigs showed high similarity to Falconiformes (28%), followed by Passeriformes (19%). These results suggest that the blood sample preparation and Illumina sequencing were successful, as the raw read assembly was undoubtedly characterized as avian. Of the P. antarcticus contigs, 2,476 and 2,450 exhibited sequence similarity to transcripts from the rock pigeon Columba livia and the peregrine falcon Falco peregrinus, respectively. In the case of the P. papua contigs, approximately 2,130 and 2,197 showed sequence similarity to transcripts from C. livia and F. peregrinus, respectively. A previous study generated a total of 15,270 and 16,070 protein-coding genes annotated from Adélie (P. adeliae) and emperor (Aptenodytes forsteri) penguin genomes [36]. However, the contigs from the current study did not match their genomic information, because only raw sequencing reads of Adélie and emperor penguins were registered in NCBI and annotation data were deposited. Thus, we analyzed orthologous similarity of the penguin transcripts after retrieving the annotated genes of Adélie and emperor penguins [36].
Ortholog comparisons showed extensive similarity within penguins (Fig. 1B). Most transcripts of each penguin matched at least one of the other species. Of the contigs, 3,969 were shared among the four penguins, whereas 1,320, 13,294, 1,699, and 9,290 contigs remained unique to the species Adélie, chinstrap, emperor, and gentoo, respectively. Within the genus Pygoscelis, 4,018 contigs were shared among the three penguins within that genus. Overall, 11,045 homologues were observed between the chinstrap and gentoo penguins and 12,387 contigs were identified as homologues between the Adélie and emperor penguins. Although a relatively small number of genes were annotated from the genomes of the Adélie and emperor penguins, this result suggests that blood transcriptomes share many genes encoding for proteins with blood-specific functions.
The specific GO composition of each principal category (i.e., cellular components, biological processes, and molecular functions) is presented for the P. antarcticus and P. papua contigs at a Level 2 percentage using default parameters. Overall, similar compositions of GO terms in each category were observed among the blood transcriptomes (Fig. 2). The vast majority of transcripts in the cellular components category were assigned to cells (34%), organelles (30%), and macromolecular complexes (12%–13%) (Fig. 2A). In terms of the biological processes category, many genes were classified as cellular (18%–19%), metabolic (15%), and single-organism processes (13%–14%) (Fig. 2B). Of the GO terms related to molecular function, many genes were categorized as binding (49%–50%) and catalytic activity (30%–31%) components (Fig. 2C). Because blood is one of the most dynamic tissues and encompasses a wide range of cellular metabolisms and numerous exogenous and endogenous factors, the diverse GO assignments analyzed by Blast2GO suggest that penguin blood performs many complex biological functions.
Of the 20 highest RPKM values, several mRNAs coding for hemoglobin genes were detected in both the P. antarcticus and P. papua transcriptomes. Because diving birds have developed elevated hemoglobin levels to hold their breath for long periods of time [37], high RPKM values for hemoglobin genes would relate to the capacity for oxygen storage and transfer during routine diving by penguins. Although only limited information is available on the correlation between abundance of hemoglobin gene and oxygen level in penguins, it was identified that hemoglobin concentration is strongly associated with free-diving physiology such as gas exchanges (e.g., oxygen storage, saturation, and depletion) in seals [38,39]. Several major vertebrate canonical signaling pathways such as the nuclear factor kappa B (NF-κB, #04064), Janus kinase/signal transducers and activators of transcription (JAK-STAT, #04630), and Toll-like receptor (TLR, #04620) are important for responding to exogenous pathogens and maintaining homeostasis [40]. In the P. antarcticus and P. papua blood transcriptomes, the most crucial proteins of innate immunity signaling were detected in the NF-κB (Fig. 3), JAK-STAT (Fig. 4A), and TLR (Fig. 4B) pathways in KEGG pathway analyses. In addition, the transcriptional involvement of immunity-related genes of the TLR, immune deficiency (Imd, #04624), transforming growth factor β (#04350), and interleukin 17 (#04657) signaling pathways were also identified. Overall, the emperor penguin had the greatest number of matched transcripts, whereas the Adélie only had a few proteins from each pathway. A relatively low amount of immunity-related genes were identified in the in the chinstrap and gentoo transcriptomes compared to the emperor penguin. In addition, there was a complete lack of several components from each pathway. This could be explained by the fact that both penguins considered in this study may not be affected by environmental factors or infections. However, the core proteins matched from each pathway suggest that the major innate immune systems are likely expressed and serve as a host defense modulator against exogenous pathological changes in penguin blood. Because the profiling of blood transcriptomes can reflect pathological changes such as immune cell circulation throughout the animal body [41-44] and penguins are sentinel species in the circumpolar region of the Southern Ocean, analyses of blood transcriptomes and hematological parameters could be used as a marker of health status, disease, and response to exposure in penguins.
Here, we presented the first whole-blood transcriptome of two penguin species, the chinstrap and gentoo, and a comparison of pathways involved in the major innate immune systems of four penguin species. Transcriptomic data were successfully obtained from the blood of the penguins and the transcriptomes covered the essential gene repertoire. These data will be useful for understanding the molecular adaptations and immune homeostasis of Antarctic penguins, thereby allowing researchers to assess their health status and resistance to diseases.
Notes
Acknowledgments
This work was supported by the Polar Genome 101 Project (Grant No. PE19080) funded by the Korea Polar Research Institute.
Fig. 1.
Major BLAST hits matching the chinstrap and gentoo blood transcripts at the phylum and species levels. Each number indicates the quantity of orthologous gene families shared by the indicated genomic database (A). Four-way Venn diagram of orthologous gene conservation in four species of penguins: Adélie (Pygoscelis adeliae), chinstrap (P. antarcticus), emperor (Aptenodytes forsteri), and gentoo (P. papua) (B). The diagram was constructed with orthologous genes identified by the best reciprocal hit method with tBLASTx (E-value < 10–10) pair-wise ortholog matches. Numbers represent unique or shared genes between penguin species.
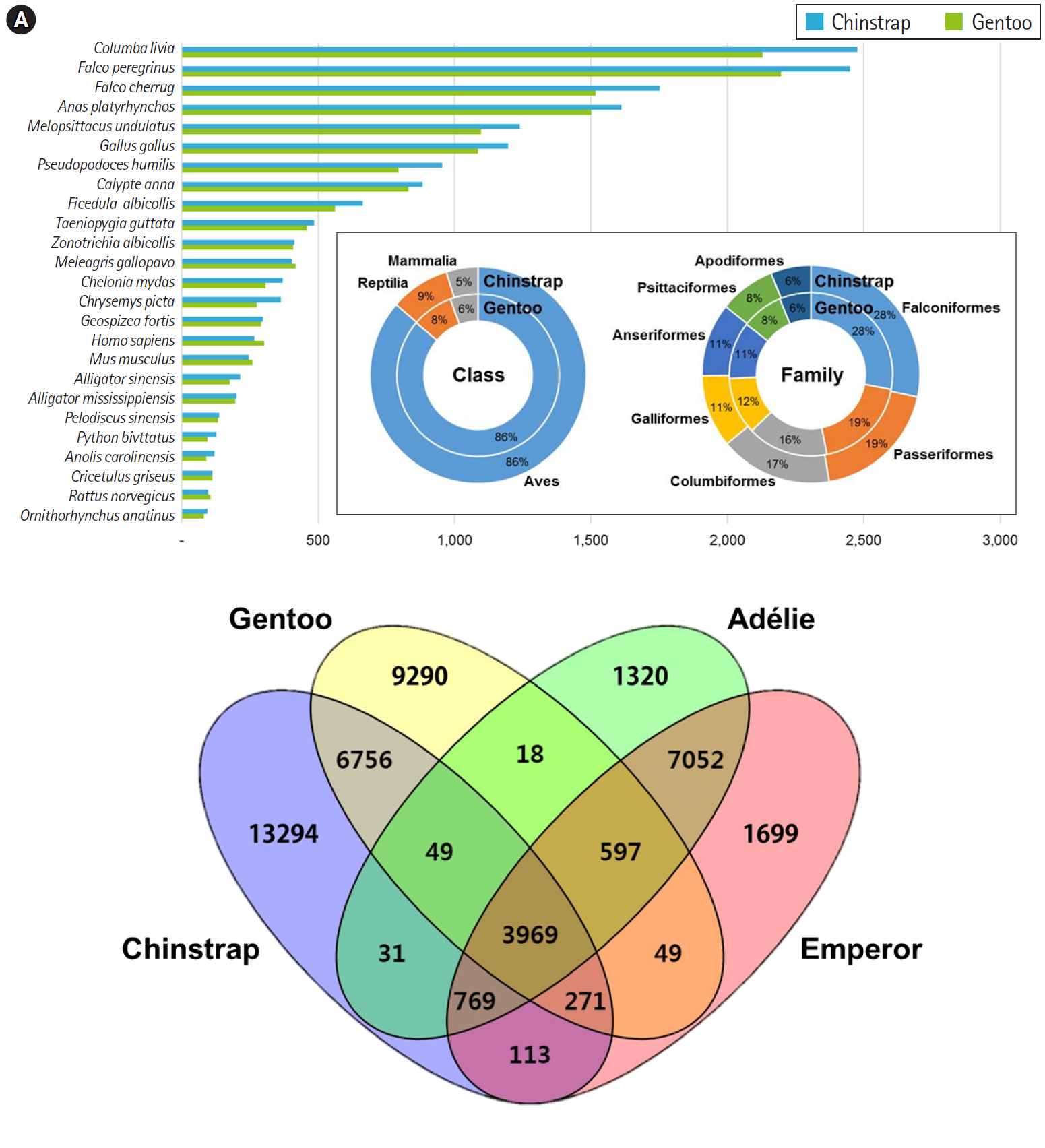
Fig. 2.
Gene Ontology analyses: cellular components (A), biological processes (B), and molecular functions (C) enriched in the chinstrap and gentoo blood transcriptomes.

Fig. 3.
Transcriptional coverage of penguin transcripts within the vertebrate nuclear factor kappa B (NF-κB) signaling pathway (#04064). Matched homologues of each penguin are highlighted in different colors.
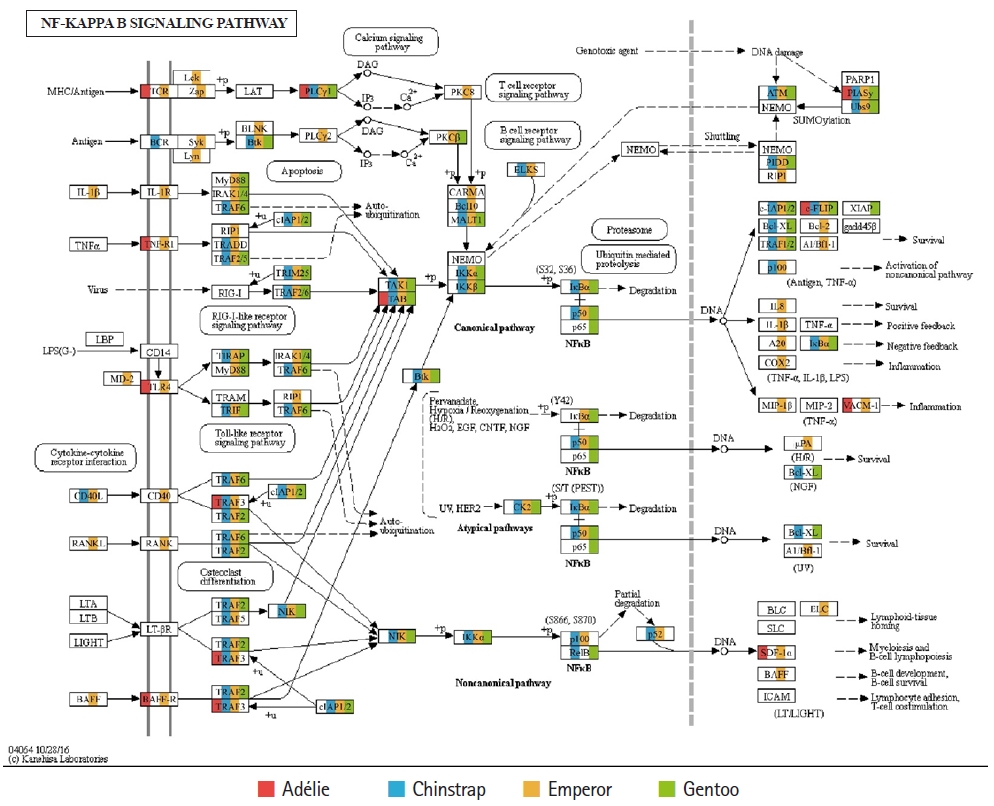
Fig. 4.
Transcriptional coverage of penguin transcripts on the vertebrate Janus kinase/signal transducers and activators of transcription (JAK-STAT) (#04630) (A) and vertebrate Toll-like receptor (TLR) signaling pathway (#04620) (B). Matched homologues of each penguin are highlighted in different colors.
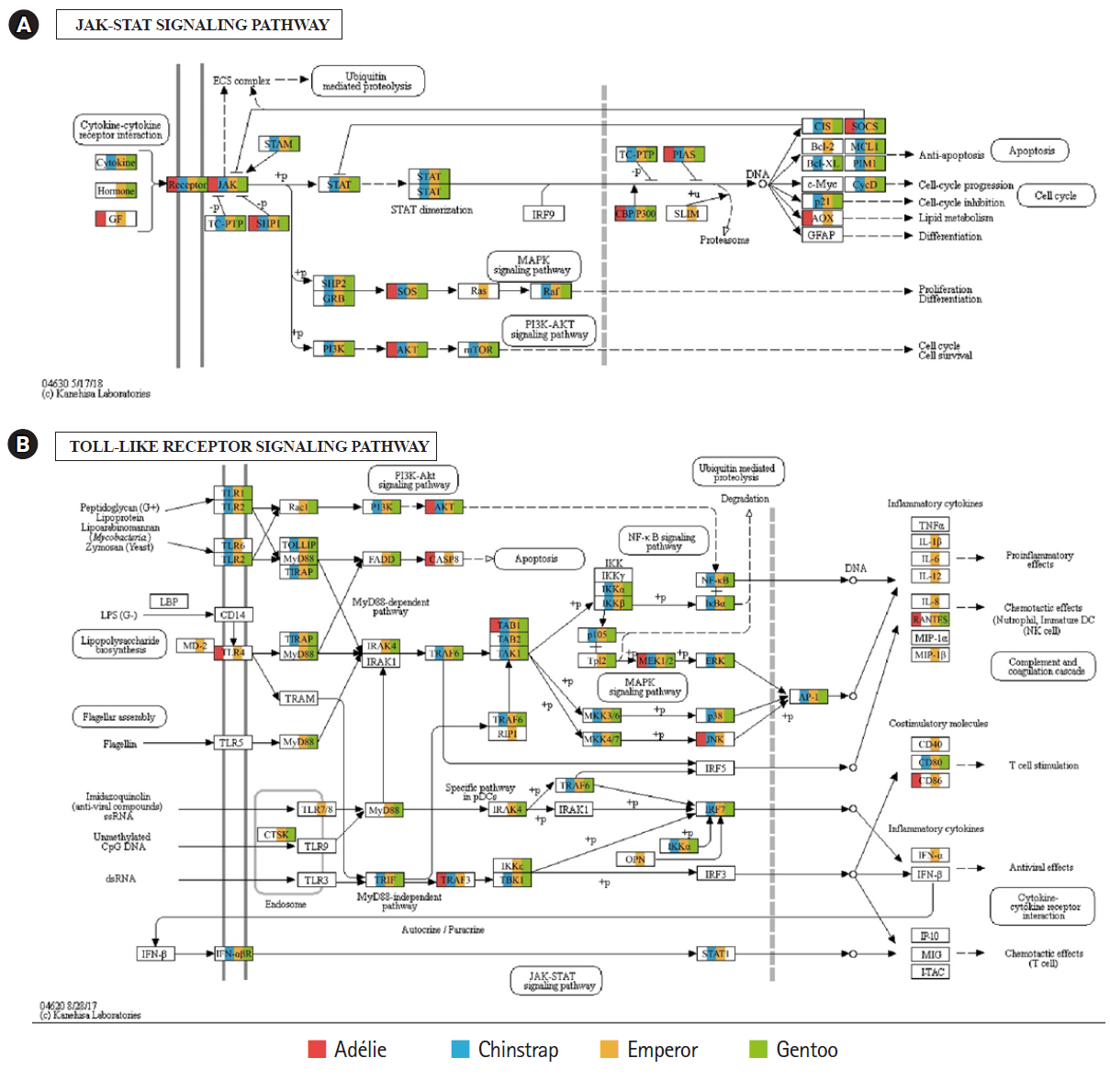
Table 1.
Characteristics of the blood transcriptomes of the chinstrap (Pygoscelis antarcticus) and gentoo (Pygoscelis papua) penguins in compliance with MIxS standards
References
1. Stonehouse B. Introduction: the Spheniscidae. In: The Biology of Penguins (Stonehouse B, ed.). London: MacMillan Press, 1975. pp. 1–15.
2. Williams TD. The Penguins: Spheniscidae. Oxford: Oxford University Press, 1995.
3. Lynnes A, Reid K, Croxall J, Trathan P. Conflict or co-existence? Foraging distribution and competition for prey between Adélie and chinstrap penguins. Mar Biol 2002;141:1165–1174.

4. Hinke JT, Salwicka K, Trivelpiece SG, Watters GM, Trivelpiece WZ. Divergent responses of Pygoscelis penguins reveal a common environmental driver. Oecologia 2007;153:845–855.


5. Negrete P, Sallaberry M, Barceló G, Maldonado K, Perona F, McGill RA, et al. Temporal variation in isotopic composition of Pygoscelis penguins at Ardley Island, Antarctic: are foraging habits impacted by environmental change? Polar Biol 2017;40:903–916.


6. Black CE. A comprehensive review of the phenology of Pygoscelis penguins. Polar Biol 2016;39:405–432.


7. Cimino MA, Lynch HJ, Saba VS, Oliver MJ. Projected asymmetric response of Adelie penguins to Antarctic climate change. Sci Rep 2016;6:28785.




9. Trathan PN, Garcia-Borboroglu P, Boersma D, Bost CA, Crawford RJ, Crossin GT, et al. Pollution, habitat loss, fishing, and climate change as critical threats to penguins. Conserv Biol 2015;29:31–41.


10. Risebrough RW, Walker W, Schmidt TT, de Lappe BW, Connors CW. Transfer of chlorinated biphenyls to Antarctica. Nature 1976;264:738–739.



11. Lenihan HS, Oliver JS, Oakden JM, Stephenson MD. Intense and localized benthic marine pollution around McMurdo station, Antarctica. Mar Pollut Bull 1990;21:422–430.

12. Klanova J, Matykiewiczova N, Macka Z, Prosek P, Laska K, Klan P. Persistent organic pollutants in soils and sediments from James Ross Island, Antarctica. Environ Pollut 2008;152:416–423.


13. Jones HI, Shellam GR. The occurrence of blood-inhabiting protozoa in captive and free-living penguins. Polar Biol 1999;21:5–10.

14. Bargagli R. Environmental contamination in Antarctic ecosystems. Sci Total Environ 2008;400:212–226.


15. Finger A, Lavers JL, Dann P, Nugegoda D, Orbell JD, Robertson B, et al. The little penguin (Eudyptula minor) as an indicator of coastal trace metal pollution. Environ Pollut 2015;205:365–377.


16. Vanstreels RE, Braga EM, Catao-Dias JL. Blood parasites of penguins: a critical review. Parasitology 2016;143:931–956.


17. Dehnhard N, Jaspers VLB, Demongin L, Van den Steen E, Covaci A, Pinxten R, et al. Organohalogenated contaminants in plasma and eggs of rockhopper penguins: does vitellogenin affect maternal transfer? Environ Pollut 2017;226:277–287.


18. Espejo W, Celis JE, GonzAlez-Acuna D, Banegas A, Barra R, Chiang G. A global overview of exposure levels and biological effects of trace elements in penguins. Rev Environ Contam Toxicol 2018;245:1–64.


19. Fantham HB, Porter A. On a Plasmodium (Plasmodium relictum var. spheniscidae, n. var.), observed in four species of penguins. Proc Zool Soc Lond 1944;114:279–292.

20. Hill AG, Howe L, Gartrell BD, Alley MR. Prevalence of Leucocytozoon spp, in the endangered yellow-eyed penguin Megadyptes antipodes. Parasitology 2010;137:1477–1485.


21. Argilla LS, Howe L, Gartrell BD, Alley MR. High prevalence of Leucocytozoon spp. in the endangered yellow-eyed penguin (Megadyptes antipodes) in the sub-Antarctic regions of New Zealand. Parasitology 2013;140:672–682.


22. Kumar KS, Kannan K, Corsolini S, Evans T, Giesy JP, Nakanishi J, et al. Polychlorinated dibenzo-p-dioxins, dibenzofurans and polychlorinated biphenyls in polar bear, penguin and south polar skua. Environ Pollut 2002;119:151–161.


23. Roosens L, Van Den Brink N, Riddle M, Blust R, Neels H, Covaci A. Penguin colonies as secondary sources of contamination with persistent organic pollutants. J Environ Monit 2007;9:822–825.


24. Schiavone A, Corsolini S, Borghesi N, Focardi S. Contamination profiles of selected PCB congeners, chlorinated pesticides, PCDD/Fs in Antarctic fur seal pups and penguin eggs. Chemosphere 2009;76:264–269.


25. Corsolini S, Borghesi N, Ademollo N, Focardi S. Chlorinated biphenyls and pesticides in migrating and resident seabirds from East and West Antarctica. Environ Int 2011;37:1329–1335.


26. Meitern R, Andreson R, Hõrak P. Profile of whole blood gene expression following immune stimulation in a wild passerine. BMC Genomics 2014;15:533.



27. Chung O, Jin S, Cho YS, Lim J, Kim H, Jho S, et al. The first whole genome and transcriptome of the cinereous vulture reveals adaptation in the gastric and immune defense systems and possible convergent evolution between the Old and New World vultures. Genome Biol 2015;16:215.



28. Tariq M, Chen R, Yuan H, Liu Y, Wu Y, Wang J, et al. De novo transcriptomic analysis of peripheral blood lymphocytes from the Chinese goose: gene discovery and immune system pathway description. PLoS One 2015;10:e0121015.



29. Videvall E, Cornwallis CK, Palinauskas V, Valkiunas G, Hellgren O. The avian transcriptome response to malaria infection. Mol Biol Evol 2015;32:1255–1267.




30. Watson H, Videvall E, Andersson MN, Isaksson C. Transcriptome analysis of a wild bird reveals physiological responses to the urban environment. Sci Rep 2017;7:44180.




31. Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014;30:2114–2120.




32. Gordon A, Hannon G. FASTQ/A short-reads pre-processing tools. FASTX-Toolkit, 2010. Accessed 2018 Dec 3. Available from: http://hannonlab.cshl.edu/fastx_toolkit.
33. Schulz MH, Zerbino DR, Vingron M, Birney E. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 2012;28:1086–1092.




34. Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005;21:3674–3676.



35. Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 2008;5:621–628.



36. Li C, Zhang Y, Li J, Kong L, Hu H, Pan H, et al. Two Antarctic penguin genomes reveal insights into their evolutionary history and molecular changes related to the Antarctic environment. Gigascience 2014;3:27.




37. Meir JU, Ponganis PJ. High-affinity hemoglobin and blood oxygen saturation in diving emperor penguins. J Exp Biol 2009;212:3330–3338.


38. Qvist J, Hill RD, Schneider RC, Falke KJ, Liggins GC, Guppy M, et al. Hemoglobin concentrations and blood gas tensions of free-diving Weddell seals. J Appl Physiol (1985) 1986;61:1560–1569.


39. Stockard TK, Levenson DH, Berg L, Fransioli JR, Baranov EA, Ponganis PJ. Blood oxygen depletion during rest-associated apneas of northern elephant seals (Mirounga angustirostris). J Exp Biol 2007;210:2607–2617.


40. Akira S, Uematsu S, Takeuchi O. Pathogen recognition and innate immunity. Cell 2006;124:783–801.


41. Mohr S, Liew CC. The peripheral-blood transcriptome: new insights into disease and risk assessment. Trends Mol Med 2007;13:422–432.


42. Chaussabel D. Assessment of immune status using blood transcriptomics and potential implications for global health. Semin Immunol 2015;27:58–66.


- TOOLS
-
METRICS

-
- 1 Crossref
- 0 Scopus
- 7,154 View
- 107 Download
- Related articles in GNI







