1. Wilkie AO, Morriss-Kay GM. Genetics of craniofacial development and malformation. Nat Rev Genet 2001;2:458–468. PMID:
11389462.


2. Schoenwolf GC, Larsen WJ. Larsen's Human Embryology. 4th ed. Philadelphia: Churchill Livingstone/Elsevier, 2009.
5. Ding YC, Wooding S, Harpending HC, Chi HC, Li HP, Fu YX,
et al. Population structure and history in East Asia. Proc Natl Acad Sci U S A 2000;97:14003–14006. PMID:
11095712.



6. Gad A, Laurino M, Maravilla KR, Matsushita M, Raskind WH. Sensorineural deafness, distinctive facial features, and abnormal cranial bones: a new variant of Waardenburg syndrome? Am J Med Genet A 2008;146A:1880–1885. PMID:
18553554.



7. Michalowicz BS, Aeppli DP, Kuba RK, Bereuter JE, Conry JP, Segal NL,
et al. A twin study of genetic variation in proportional radiographic alveolar bone height. J Dent Res 1991;70:1431–1435. PMID:
1960253.


8. Carels C, Van Cauwenberghe N, Savoye I, Willems G, Loos R, Derom C,
et al. A quantitative genetic study of cephalometric variables in twins. Clin Orthod Res 2001;4:130–140. PMID:
11553097.


9. Naini FB, Moss JP. Three-dimensional assessment of the relative contribution of genetics and environment to various facial parameters with the twin method. Am J Orthod Dentofacial Orthop 2004;126:655–665. PMID:
15592212.


10. Amini F, Borzabadi-Farahani A. Heritability of dental and skeletal cephalometric variables in monozygous and dizygous Iranian twins. Orthod Waves 2009;68:72–79.

11. Baydaş B, Erdem A, Yavuz I, Ceylan I. Heritability of facial proportions and soft-tissue profile characteristics in Turkish Anatolian siblings. Am J Orthod Dentofacial Orthop 2007;131:504–509. PMID:
17418717.


12. Susanne C. Genetic and environmental influences on morphological characteristics. Ann Hum Biol 1975;2:279–287. PMID:
16431681.


13. Kohn LA. The role of genetics in craniofacial morphology and growth. Annu Rev Anthropol 1991;20:261–278.

14. Ermakov S, Kobyliansky E, Livshits G. Quantitative genetic study of head size related phenotypes in ethnically homogeneous Chuvasha pedigrees. Ann Hum Biol 2005;32:585–598. PMID:
16316915.


15. Johannsdottir B, Thorarinsson F, Thordarson A, Magnusson TE. Heritability of craniofacial characteristics between parents and offspring estimated from lateral cephalograms. Am J Orthod Dentofacial Orthop 2005;127:200–207. PMID:
15750539.


16. Martínez-Abadías N, Esparza M, Sjøvold T, González-José R, Santos M, Hernández M. Heritability of human cranial dimensions: comparing the evolvability of different cranial regions. J Anat 2009;214:19–35. PMID:
19166470.



17. Alkhudhairi TD, Alkofide EA. Cephalometric craniofacial features in Saudi parents and their offspring. Angle Orthod 2010;80:1010–1017. PMID:
20677948.



18. Karmakar B, Ermakov S, Yakovenko K, Kobyliansky E. Genetic determination of head-size-related anthropometric traits in an ethnically homogeneous sample of 373 Indian pedigrees of West Bengal. Hum Biol 2007;79:501–514. PMID:
18478966.


19. Jelenkovic A, Poveda A, Susanne C, Rebato E. Contribution of genetics and environment to craniofacial anthropometric phenotypes in Belgian nuclear families. Hum Biol 2008;80:637–654. PMID:
19728541.


20. Zhang X, Hans MG, Graham G, Kirchner HL, Redline S. Correlations between cephalometric and facial photographic measurements of craniofacial form. Am J Orthod Dentofacial Orthop 2007;131:67–71. PMID:
17208108.


21. Farkas LG. Anthropometry of the Head and Face. New York: Raven Press, 1994.
22. Sharma S. Applied Multivariate Techniques. New York: John Wiley and Sons Inc., 1996.
23. Cheung WY, Le LW, Zimmermann C. Symptom clusters in patients with advanced cancers. Support Care Cancer 2009;17:1223–1230. PMID:
19184126.


24. Almasy L, Blangero J. Multipoint quantitative-trait linkage analysis in general pedigrees. Am J Hum Genet 1998;62:1198–1211. PMID:
9545414.



25. Keen KJ, Elston RC. Robust asymptotic sampling theory for correlations in pedigrees. Stat Med 2003;22:3229–3247. PMID:
14518025.


26. Ferrario VF, Sforza C, Miani A, Tartaglia G. Craniofacial morphometry by photographic evaluations. Am J Orthod Dentofacial Orthop 1993;103:327–337. PMID:
8480698.


27. Weinberg SM, Scott NM, Neiswanger K, Brandon CA, Marazita ML. Digital three-dimensional photogrammetry: evaluation of anthropometric precision and accuracy using a Genex 3D camera system. Cleft Palate Craniofac J 2004;41:507–518. PMID:
15352857.


28. Nagle E, Teibe U, Kapoka D. Craniofacial anthropometry in a group of healthy Latvian residents. Acta Med Litu 2005;12:47–53.
29. Arya R, Duggirala R, Comuzzie AG, Puppala S, Modem S, Busi BR,
et al. Heritability of anthropometric phenotypes in caste populations of Visakhapatnam, India. Hum Biol 2002;74:325–344. PMID:
12180759.


30. Raposo-do-Amaral CM, Krieger H, Cabello PH, Beiguelman B. Heritability of quantitative orbital traits. Hum Biol 1989;61:551–557. PMID:
2591913.

31. Im SW, Kim HJ, Lee MK, Yi JH, Jargal G, Sung J,
et al. Genome-wide linkage analysis for ocular and nasal anthropometric traits in a Mongolian population. Exp Mol Med 2010;42:799–804. PMID:
21150245.



32. Manfredi C, Martina R, Grossi GB, Giuliani M. Heritability of 39 orthodontic cephalometric parameters on MZ, DZ twins and MN-paired singletons. Am J Orthod Dentofacial Orthop 1997;111:44–51. PMID:
9009923.


33. Hauspie RC, Susanne C, Defrise-Gussenhoven E. Testing for the presence of genetic variance in factors of face measurements of Belgian twins. Ann Hum Biol 1985;12:429–440. PMID:
4062238.


34. Hopper JL, Bishop DT, Easton DF. Population-based family studies in genetic epidemiology. Lancet 2005;366:1397–1406. PMID:
16226618.


35. Borecki IB, Province MA. Genetic and genomic discovery using family studies. Circulation 2008;118:1057–1063. PMID:
18765388.


36. Fuller SJ, Papaemmanuil E, McKinnon L, Webb E, Sellick GS, Dao-Ung LP,
et al. Analysis of a large multi-generational family provides insight into the genetics of chronic lymphocytic leukemia. Br J Haematol 2008;142:238–245. PMID:
18503587.


37. Allanson JE. Objective techniques for craniofacial assessment: what are the choices? Am J Med Genet 1997;70:1–5. PMID:
9129732.


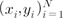 and arbitrary weights {wi} [25].
and arbitrary weights {wi} [25].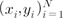 and arbitrary weights {wi} [25].
and arbitrary weights {wi} [25].