 |
 |
- Search
| Genomics Inform > Volume 14(4); 2016 > Article |
Abstract
Osteoporotic fractures (OFs) are critical hard outcomes of osteoporosis and are characterized by decreased bone strength induced by low bone density and microarchitectural deterioration in bone tissue. Most OFs cause acute pain, hospitalization, immobilization, and slow recovery in patients and are associated with increased mortality. A variety of genetic studies have suggested associations of genetic variants with the risk of OF. Genome-wide association studies have reported various single-nucleotide polymorphisms and copy number variations (CNVs) in European and Asian populations. To identify CNV regions associated with OF risk, we conducted a genome-wide CNV study in a Korean population. We performed logistic regression analyses in 1,537 Korean subjects (299 OF cases and 1,238 healthy controls) and identified a total of 8 CNV regions significantly associated with OF (p < 0.05). Then, one CNV region located on chromosome 20q13.12 was selected for experimental validation. The selected CNV region was experimentally validated by quantitative polymerase chain reaction. The CNV region of chromosome 20q13.12 is positioned upstream of a family of long non-coding RNAs, LINC01260. Our findings could provide new information on the genetic factors associated with the risk of OF.
Osteoporotic fractures (OFs) result from a decrease in bone strength, which can be induced by low bone mass and microarchitectural deterioration in bone tissue [1]. OFs are the critical hard outcome of osteoporosis, a disease that affects more than 75 million people in the United States, Europe, and Japan. With a lifetime fracture risk of 30% to 40% (vertebral or non-vertebral fractures), osteoporosis has an incidence rate similar to that of coronary artery disease. Furthermore, with the exception of forearm fractures, OFs are associated with increased mortality. Most fractures cause acute pain and lead to patient hospitalization, immobilization, and slow recovery [2,3,4].
Genetic studies have suggested a correlation of genetic variations with OF and its related traits. First, a candidate gene study has reported that common non-synonymous variants in low-density lipoprotein receptor-related protein 5 (LRP5) are consistently associated with bone mineral density (BMD) and fracture risk [5]. Furthermore, meta-analyses of genome-wide association studies (GWASs) have identified single-nucleotide polymorphisms (SNPs) located in more than 56 loci independently associated with BMD, and some of these studies have also found associations with fracture risk [6,7,8,9,10,11,12,13].
Copy number variations (CNVs) also have shown associations with OF. A genome-wide CNV study performed in a Chinese population identified a deletion variant of UDP glucuronosyltransferase 2 family, polypeptide B17 (UGT2B17) in chromosome 4q13.2 associated with OF [14]; however, this variant was not replicated in a study of Caucasian women [15], possibly showing ethnic specificity. Additionally, a rare deletion variant located on chromosome 6p25.1 showed an association with the risk of OF in a Dutch population [16].
Particularly, chromosome 20q13.12 is known as an enriched region of histone modifications. Many genes located on chromosome 20q13.12 have enriched levels of histone modifications, such as di-methylated lysine 4 of histone 3 (H3K4me2) and tri-methylated lysine 4 of histone 3 (H3K4me3), in their promoter regions [17]. H3K4me2 and H3K4me3 regulate the expression of the Runx2 gene, which encodes a transcription factor controlling bone development and osteoblast differentiation [18]. Although there is no previous study on an association between chromosome 20q13.12 and bone-related traits, an SNP in another 20q region (rs4811196 in CTNBL1, located on 20q11.23) has been reported to be associated with BMD [19].
In this study, we conducted an association study in the Korean population to identify new susceptibility loci for the risk of OF. We identified 8 CNV regions associated with the risk of OF through a statistical analysis. Among the 8 regions, we selected and validated the existence of one CNV by quantitative PCR.
A total of 10,038 participants from the Korea Association Resource (KARE) were recruited. Among the recruited individuals, 1,537 subjects, consisting of 299 OF cases and 1,238 normal controls, who agreed to an X-ray examination were genotyped with the NimbleGen HD2 3 ├Ś 720K comparative genomic hybridization array (aCGH). This study was approved by the Institutional Review Board of the Korea Centers for Disease Control and Prevention, and written informed consent was provided to all participants, including cases and controls. The clinical characteristics of the study participants are summarized in Table 1. We used low trauma fracture events to distinguish between fracture and non-fracture groups of the KARE study. Fractures were only included if they had occurred with low trauma (e.g., fall from standing height or less) at any of six sites (hip, wrist, humerus, rib, pelvis, and vertebra) after the age of 40 years. Fractures clearly caused by high trauma events (e.g., motor vehicle accidents, violence, or falls from more than the standing height of the individual) were excluded.
We used the Roche NimbleGen 3 ├Ś 720 K aCGH platform for CNV calling. This platform was designed as a whole-genome and CNV-focused type, composed of a total of 720,000 probes. The median inter-probe spacing of the backbone was <5 kb. DNA extracted from the NA10851 cell line was utilized as a reference for the aCGH to yield the signal intensity ratio with hg18/NCBI build 36. All of the samples in our study satisfied experimental control metrics, such as chromosome X shift and mad.1dr, with NimbleScan v.2.5 to adapt systemic biases in the microarray experiment. The signal intensity ratio of each probe was converted into log2 scale with the positions of the probes after a quality control process.
CNV discovery consisted of two steps: CNV calling and determination of the CNV region. For CNV calling, we used a package, called ŌĆ£Genome Alteration Detection Analysis (GADA),ŌĆØ which implements a flexible and efficient pipeline to detect copy number alterations from microarray data [20]. To avoid a limitation caused in the single detection tool, we found the best parameter (T = 10, alpha = 0.2, and Min-SegLen = 10) that compared a known CNV region and CNVs tested using several parameters [21]. CNV was employed as the average log2 ratio of ┬▒0.25 of probes in all individual. To estimate CNV genotypes, a CNV region was defined with a log2 ratio identified between the reference sample and test sample in the calling process. It is hard to define exact genotype cluster of a CNV detected in a single individual. To discover CNV regions, we employed CNVs that were called in over three individuals [22].
CNV genotypes consist of zero copies, one copy, and two copies. To do this, we used an R package, named ŌĆśCNVtools,ŌĆÖ with default parameters to estimate CNV genotypes [23]. CNVtools is a mixture model for one-dimensional CNV data summary that can separate samples into each CNV genotype. To estimate genotype, we employed the linear discrimination function, which uses a raw signal and pca signal calculated from the average log2 ratio of all individuals using CNVtools. CNV genotypes consist of three genotype groups: ŌĆ£single-class,ŌĆØ ŌĆ£multiple-class,ŌĆØ and ŌĆ£ambiguousŌĆØ (Supplementary Fig. 1) [24,25]. We performed an association study with only well-clustered ŌĆ£multiple-classŌĆØ CNV regions.
Logistic regression analysis was performed using R package version 3.0.2 to find significant association signals between CNV loci and OF risk. Age, height, and weight were adjusted as covariates to calibrate the statistical significance. Genotypes of each CNV were coded as 0 (homozygous deletion) and 1 (heterozygous deletion) for two-class CNV loci. In the case of three-class CNVs, their genotypes were coded as 0 (homozygous deletion), 1 (heterozygous deletion), and 2 (normal).
We carried out quantitative PCR using the TaqMan Copy Number Assay (Life Technologies, Foster City, CA, USA) according to the manufacturer's protocols to verify estimated CNV regions. In total, seven pre-designed and one custom designed probe was used to validate the existence of the CNV. Validation samples, including cases and controls, were randomly selected from each estimated genotype cluster. All validation experiments were replicated 3 times to increase the accuracy of the validation. The copy number in each individual was calculated with Copy Caller v2.0 using the comparative threshold cycle (CT) method according to the manufacturer's protocols.
A total of 1,537 Korean individuals, consisting of 299 OF patients and 1,238 normal controls, participated in this study (Table 1). There were 415 males and 1,122 females in the overall population. The ratio of females in controls was slightly higher than in the cases (70% in cases vs. 72% in controls, data not shown). Mean age, height, weight, and body mass index showed no significant differences between cases and controls.
To select candidate CNV regions, we manually performed a visual inspection to classify well-clustered ŌĆ£multiple-classŌĆØ regions into genotype groups. In total, 3,660 multiple-class CNV regions were selected for the association study. Of the 3,660 CNV regions, 518 CNV regions were two-class regions (one and two copies) and 3,142 were three-class regions (zero, one, and two copies) (Supplementary Fig. 1).
We performed logistic regression analysis to evaluate associations between identified CNV regions and OF risk. As a result, we identified a total of eight CNV loci associated with OF (p < 0.05) (Table 2)ŌĆödistributed on chromosome 4q13.1, 4q13.2, 9q34.2, 13q12.2, 14q24.3, 14q31.3, 20p13, and 20q13.12. The OF-associated loci were located in intergenic regions near SRIP1; TMPRSS11E; POLR1D; LOC-100421611; a family of long non-coding RNAs (lncRNA), LINC01260; and genic regions of ABO, ELMSAN1, and SIRPA. The base-pair lengths of the eight regions ranged from 1.9 kb to 18.6 kb. The CNVs in ABO and SIRPA included both exonic and intronic regions of each gene, whereas the CNV locus in ELMSAN1 included only intronic regions.
We selected one CNV region located on chromosome 20q13.12ŌĆöchr20: 42739446ŌĆō42741539ŌĆöor validation based on position and expected biological function. The allele frequencies of chr20: 42739446ŌĆō42741539 are indicated in Supplementary Table 1 and Supplementary Fig. 2. First, we conducted in silico verification for the existence of the CNV region using web-based public data. Many previously reported CNVs, including the chr20: 42739446ŌĆō42741539 region, were identified using the UCSC Genome Browser (http://genome.uscs.edu/cgi-bin/hgGateway) (Supplementary Fig. 3). Also, frequencies of reported CNVs evaluated in various ethnic groups were presented in the Database of Genomic Variants (DGV, http://dgv.tcag.ca/dgv/app/home) (Supplementary Table 1).
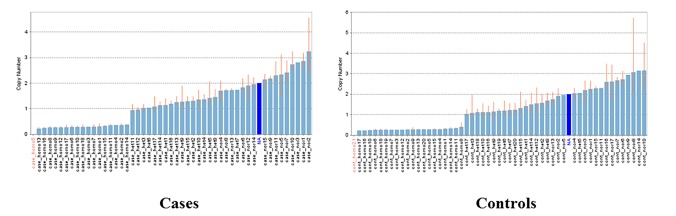
To experimentally verify the accuracy of the CNV, we performed quantitative PCR. The position of the CNV probe was defined based on hg19/NCBI build 37. The probe ID used for the TaqMan genotyping assay of the validated CNVs is listed in Table 3. Applications of a positive predictive value as a measurement standard of accuracy indicated the proportion of positive results in the validation of the CNV region. The positive predictive value of the verified CNV region evaluated in our validation was 0.82. Fig. 1 shows the genotype of the identified CNV region, verified by quantitative PCR.
In this study, we performed a GWAS to investigate the associations between identified CNV regions and the risk of OF. A total of eight CNV loci were identified to be nominally associated with OF. For validation, we selected one CNV region based on position and expected biological meaning among the eight loci. The CNV, which is located on chromosome 20q13.12 (intergenic region near LINC01260) was experimentally validated through quantitative PCR.
An lncRNA is defined as a non-coding transcript having a length of more than 200 nucleotides [26] and is considered ŌĆ£junkŌĆØ that is non-functional initially [27]. However, the functional roles of lncRNAs have now been identified, such as key regulators of transcriptional and translational products that affect cell functions [28,29,30,31]. lncRNAs play a role in epigenetic effects, such as histone modification (e.g., methylation), which regulates gene expression [32]. A significant histone modification, monomethylation of lysine 4 on histone H3 (H3K4me1) in human mammary epithelial cells, was predicted in our identified CNV region near an lncRNA, LINC01260, using University of California Santa Cruz (UCSC) Genome Browser (http://genome.uscs.edu/cgi-bin/hgGateway) (Supplementary Fig. 4). A previous genome-wide study has identified several runs of homozygosity (ROHs) associated with BMD and observed the enrichment of H3K4me1 in two ROHs, ROH15q22.3 and ROH1p31.1, by an analysis of histone modification marks in the Gm12878 cell line [33]. Taken together, our identified CNV region on 20q13.12 might have a putative effect on the risk of OF by histone modification of the promoter region of LINC01260.
In addition, a CNV on chromosome 20q13.12 is located in a region about 35 kb apart from the WISP2 gene. The WISP2 gene has some evidence of functions associated with bone-related traits. WISP2, also known as CCN family member 5 (CCN5) or connective tissue growth factor like (CTGF-L), was identified as a member of the CCN family, downstream of the WNTŌĆō╬▓-catenin signaling pathway, using subtractive hybridization between WNT1-transformed mammary epithelial cells and parental cells [34]. The WNTŌĆō╬▓-catenin signaling pathway has been suggested to be a major component in the accrual, regulation, and maintenance of bone mass [35]. WISP2 consists of 3 domains: an insulin-like growth factor binding domain, a von Willebrand Factor type C motif, and a thrombospondin type I repeat. The mRNA of WISP2 is expressed in primary human osteoblasts, fibroblasts, ovary, testes, and heart. WISP2 protein is secreted from primary human osteoblasts and fibroblasts [36]. A previous study has shown the relatedness of increased mRNA expression of WISP2 with rheumatoid arthritis and osteoarthritis [34]. Furthermore, the WISP2 gene has been reported as a candidate gene that is associated with an OF-related trait, BMD [37].
WISP2 expression is affected by the action of LRP5. LRP5 has been known to play a role in signaling through WNTŌĆō╬▓-catenin and acts as a co-receptor with the frizzled family of receptors [38]. Previous research has reported that the LRP5 mutation (G171V), inducing the enhancement of structural strength and material properties of skeleton and bone mass, is correlated with increased expression of WISP2 [38]. Also, WISP2 is suggested to be involved in the frizzled receptors/LRP5/6 pathway by phosphorylation of LRP5 [39]. Considering the previously reported functions and interactions with LRP5 in the WNTŌĆō╬▓-catenin signaling pathway, WISP2 could be an important component for bone formation or strength, which might affect the risk of OF.
In conclusion, we identified a total of 8 CNV regions associated with OF in the Korean population. Among the 8 CNVs, one locus, located on chromosome 20q13.12, was selected for validation and verified by quantitative PCR. Although further study in a larger number of study subjects is needed, to our knowledge, the current report is the first to investigate the association with the risk of OF. Our findings from this study could provide new insights into the genetic factors associated with OF risk.
Acknowledgments
This work was supported by intramural grants from the Korea National Institute of Health (2012-N73004-00 & 2012-N73005-00) and was provided with biospecimen data from the Korea Genome Analysis Project (4845-301), the Korean Genome and Epidemiology Study (4851-302), and the Korea Biobank Project (4851-307).
Supplementary materials
Supplementary data including one table and four figures can be found with this article online at http://www.genominfo.org/src/sm/gni-14-216-s001.pdf.
Supplementary Table 1
Comparison between frequencies of previously reported CNVs in the DGV database and the identified CNV in our study
Supplementary Fig. 1
Classes of copy number variation (CNV). We assigned individuals to each CNV cluster according to the log2 ratio between the test sample and reference sample. (A) Single-class CNVs, in which all individuals of the CNV region belonged to one cluster. (B) Two-class CNVs. (C) Three-class CNVs. (D) CNVs that were separated with ambiguous clusters. Only multi-class (B and C) clusters were used for the association analysis.
Supplementary Fig. 2
Scatter plot of the copy number variation (CNV) region, chr20: 42739446ŌĆō42741539. Light green, purple, and blue dots indicate normal copy, one-copy deletion, and two-copy deletion, respectively. Linear discriminant function (LDF) values from CNVtools were used for the clustering analysis. SegMean is segmental mean value (log R ratio). A loss is indicated in blue and purple.
Supplementary Fig. 3
Screenshot of genomic region chr20: 42739446ŌĆō42741539 from University of California Santa Cruz (UCSC) Genome Browser obtained from Database of Genomic Variants (DGV).
Supplementary Fig. 4
Copy number variation (CNV) at chromosome 20q13.12 and its nearby genes. A light blue-filled area indicates the identified CNV region in this study. The base pair position of the CNV region was calculated based on hg18/NCBI build 36 by the University of California Santa Cruz (UCSC) Genome Browser. Genes located near the CNV region and H3K4me1 signals (green peaks) of a cell line, human mammary epithelial cell, in this region are described.
References
1. Kanis JA, McCloskey EV, Johansson H, Cooper C, Rizzoli R, Reginster JY, et al. European guidance for the diagnosis and management of osteoporosis in postmenopausal women. Osteoporos Int 2013;24:23ŌĆō57. PMID: 23079689.



2. Hernlund E, Svedbom A, Iverg├źrd M, Compston J, Cooper C, Stenmark J, et al. Osteoporosis in the European Union: medical management, epidemiology and economic burden. A report prepared in collaboration with the International Osteoporosis Foundation (IOF) and the European Federation of Pharmaceutical Industry Associations (EFPIA). Arch Osteoporos 2013;8:136. PMID: 24113837.



4. Harvey N, Dennison E, Cooper C. Osteoporosis: impact on health and economics. Nat Rev Rheumatol 2010;6:99ŌĆō105. PMID: 20125177.


5. van Meurs JB, Trikalinos TA, Ralston SH, Balcells S, Brandi ML, Brixen K, et al. Large-scale analysis of association between LRP5 and LRP6 variants and osteoporosis. JAMA 2008;299:1277ŌĆō1290. PMID: 18349089.



6. Duncan EL, Danoy P, Kemp JP, Leo PJ, McCloskey E, Nicholson GC, et al. Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet 2011;7:e1001372. PMID: 21533022.



7. Hsu YH, Zillikens MC, Wilson SG, Farber CR, Demissie S, Soranzo N, et al. An integration of genome-wide association study and gene expression profiling to prioritize the discovery of novel susceptibility loci for osteoporosis-related traits. PLoS Genet 2010;6:e1000977. PMID: 20548944.



8. Kung AW, Xiao SM, Cherny S, Li GH, Gao Y, Tso G, et al. Association of JAG1 with bone mineral density and osteoporotic fractures: a genome-wide association study and follow-up replication studies. Am J Hum Genet 2010;86:229ŌĆō239. PMID: 20096396.



9. Richards JB, Rivadeneira F, Inouye M, Pastinen TM, Soranzo N, Wilson SG, et al. Bone mineral density, osteoporosis, and osteoporotic fractures: a genome-wide association study. Lancet 2008;371:1505ŌĆō1512. PMID: 18455228.



10. Rivadeneira F, Styrk├Īrsdottir U, Estrada K, Halld├│rsson BV, Hsu YH, Richards JB, et al. Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat Genet 2009;41:1199ŌĆō1206. PMID: 19801982.



11. Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. Multiple genetic loci for bone mineral density and fractures. N Engl J Med 2008;358:2355ŌĆō2365. PMID: 18445777.


12. Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. New sequence variants associated with bone mineral density. Nat Genet 2009;41:15ŌĆō17. PMID: 19079262.


13. Estrada K, Styrkarsdottir U, Evangelou E, Hsu YH, Duncan EL, Ntzani EE, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet 2012;44:491ŌĆō501. PMID: 22504420.



14. Yang TL, Chen XD, Guo Y, Lei SF, Wang JT, Zhou Q, et al. Genome-wide copy-number-variation study identified a susceptibility gene, UGT2B17, for osteoporosis. Am J Hum Genet 2008;83:663ŌĆō674. PMID: 18992858.



15. Chew S, Mullin BH, Lewis JR, Spector TD, Prince RL, Wilson SG. Homozygous deletion of the UGT2B17 gene is not associated with osteoporosis risk in elderly Caucasian women. Osteoporos Int 2011;22:1981ŌĆō1986. PMID: 20878390.



16. Oei L, Hsu YH, Styrkarsdottir U, Eussen BH, de Klein A, Peters MJ, et al. A genome-wide copy number association study of osteoporotic fractures points to the 6p25.1 locus. J Med Genet 2014;51:122ŌĆō131. PMID: 24343915.


17. Akan P, Sahl├®n M, Deloukas P. A histone map of human chromosome 20q13.12. PLoS One 2009;4:e4479. PMID: 19229332.



18. Rojas A, Aguilar R, Henriquez B, Lian JB, Stein JL, Stein GS, et al. Epigenetic control of the bone-master Runx2 gene during osteoblast-lineage commitment by the histone demethylase JARID1B/KDM5B. J Biol Chem 2015;290:28329ŌĆō28342. PMID: 26453309.



19. Kiel DP, Demissie S, Dupuis J, Lunetta KL, Murabito JM, Karasik D. Genome-wide association with bone mass and geometry in the Framingham Heart Study. BMC Med Genet 2007;8(Suppl 1):S14. PMID: 17903296.



20. Pique-Regi R, C├Īceres A, Gonz├Īlez JR. R-Gada: a fast and flexible pipeline for copy number analysis in association studies. BMC Bioinformatics 2010;11:380. PMID: 20637081.



21. Pique-Regi R, Monso-Varona J, Ortega A, Seeger RC, Triche TJ, Asgharzadeh S. Sparse representation and Bayesian detection of genome copy number alterations from microarray data. Bioinformatics 2008;24:309ŌĆō318. PMID: 18203770.



22. Moon S, Keam B, Hwang MY, Lee Y, Park S, Oh JH, et al. A genome-wide association study of copy-number variation identifies putative loci associated with osteoarthritis in Koreans. BMC Musculoskelet Disord 2015;16:76. PMID: 25880085.




23. Barnes C, Plagnol V, Fitzgerald T, Redon R, Marchini J, Clayton D, et al. A robust statistical method for case-control association testing with copy number variation. Nat Genet 2008;40:1245ŌĆō1252. PMID: 18776912.



24. Wellcome Trust Case Control Consortium. Craddock N, Hurles ME, Cardin N, Pearson RD, Plagnol V, et al. Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 2010;464:713ŌĆō720. PMID: 20360734.



25. Kim YK, Moon S, Hwang MY, Kim DJ, Oh JH, Kim YJ, et al. Gene-based copy number variation study reveals a microdeletion at 12q24 that influences height in the Korean population. Genomics 2013;101:134ŌĆō138. PMID: 23147675.


26. Frith MC, Bailey TL, Kasukawa T, Mignone F, Kummerfeld SK, Madera M, et al. Discrimination of non-protein-coding transcripts from protein-coding mRNA. RNA Biol 2006;3:40ŌĆō48. PMID: 17114936.


27. van Bakel H, Nislow C, Blencowe BJ, Hughes TR. Most ŌĆ£dark matterŌĆØ transcripts are associated with known genes. PLoS Biol 2010;8:e1000371. PMID: 20502517.



28. Mercer TR, Qureshi IA, Gokhan S, Dinger ME, Li G, Mattick JS, et al. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci 2010;11:14. PMID: 20137068.



29. Chen LL, Carmichael GG. Decoding the function of nuclear long non-coding RNAs. Curr Opin Cell Biol 2010;22:357ŌĆō364. PMID: 20356723.



30. Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB, et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 2008;18:1433ŌĆō1445. PMID: 18562676.



31. Loewer S, Cabili MN, Guttman M, Loh YH, Thomas K, Park IH, et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet 2010;42:1113ŌĆō1117. PMID: 21057500.



32. Cao J. The functional role of long non-coding RNAs and epigenetics. Biol Proced Online 2014;16:11. PMID: 25276098.



33. Yang TL, Guo Y, Zhang JG, Xu C, Tian Q, Deng HW. Genome-wide survey of runs of homozygosity identifies recessive loci for bone mineral density in Caucasian and Chinese populations. J Bone Miner Res 2015;30:2119ŌĆō2126. PMID: 25983029.



34. Tanaka I, Morikawa M, Okuse T, Shirakawa M, Imai K. Expression and regulation of WISP2 in rheumatoid arthritic synovium. Biochem Biophys Res Commun 2005;334:973ŌĆō978. PMID: 16038875.


35. Gong Y, Slee RB, Fukai N, Rawadi G, Roman-Roman S, Reginato AM, et al. LDL receptor-related protein 5 (LRP5) affects bone accrual and eye development. Cell 2001;107:513ŌĆō523. PMID: 11719191.


36. Kumar S, Hand AT, Connor JR, Dodds RA, Ryan PJ, Trill JJ, et al. Identification and cloning of a connective tissue growth factor-like cDNA from human osteoblasts encoding a novel regulator of osteoblast functions. J Biol Chem 1999;274:17123ŌĆō17131. PMID: 10358067.


37. Yerges LM, Klei L, Cauley JA, Roeder K, Kammerer CM, Moffett SP, et al. High-density association study of 383 candidate genes for volumetric BMD at the femoral neck and lumbar spine among older men. J Bone Miner Res 2009;24:2039ŌĆō2049. PMID: 19453261.



38. Robinson JA, Chatterjee-Kishore M, Yaworsky PJ, Cullen DM, Zhao W, Li C, et al. Wnt/beta-catenin signaling is a normal physiological response to mechanical loading in bone. J Biol Chem 2006;281:31720ŌĆō31728. PMID: 16908522.


39. Hammarstedt A, Hedjazifar S, Jenndahl L, Gogg S, Gr├╝nberg J, Gustafson B, et al. WISP2 regulates preadipocyte commitment and PPARgamma activation by BMP4. Proc Natl Acad Sci U S A 2013;110:2563ŌĆō2568. PMID: 23359679.



Fig.┬Ā1
Validated genotypes of a 20q13.12 copy number variation (CNV) region in this study. Genotypes of a CNV region on chromosome 20q13.12. The copy number state of cases (left) and control samples (right). The higher bar, lower bar, and no bar in each figure represent a normal number of copies, heterozygous deletion, and homozygous deletion, respectively. The blue bar means the copy number state of the NA10851 sample, which was used as the reference sample.
Table┬Ā1.
Clinical characteristics of study subjects
Table┬Ā2.
Association results of 8 CNV regions with risk of OF (p < 0.05)
| Chr | Starta | Enda | Length (bp) | Gene | p-value |
|---|---|---|---|---|---|
| 4 | 59662401 | 59670658 | 8,258 | Intergenic (nearby SRIP1) | 0.0001 |
| 4 | 69056562 | 69075975 | 19,414 | Intergenic (nearby TMPRSS11E) | 0.0084 |
| 9 | 135120477 | 135122444 | 1,968 | ABO (intron and exon) | 0.0085 |
| 13 | 27153540 | 27157992 | 4,453 | Intergenic (nearby POLR1D) | 0.003 |
| 14 | 73310052 | 73320816 | 10,765 | ELMSAN1 (intron) | 0.0054 |
| 14 | 84366904 | 84378906 | 12,003 | Intergenic (nearby LOC100421611) | 0.0078 |
| 20 | 1843096 | 1861664 | 18,569 | SIRPA (intron and exon) | 0.0058 |
| 20 | 42739446 | 42741539 | 2,094 | Intergenic (nearby LINC01260) | 0.0016 |
Table┬Ā3.
Results of CNV validation though TaqMan copy number assays
| Chr | Start | End | Probe ID | CNV genotype | TP | FP | PPVa |
|---|---|---|---|---|---|---|---|
| 20 | 43306032 | 43308125 | Hs07172967_cn | 0/1/2 | 84 | 18 | 0.82 |
- TOOLS
-
METRICS

-
- 3 Crossref
- 0 Scopus
- 6,062 View
- 92 Download
- Related articles in GNI





