 |
 |
- Search
| Genomics Inform > Volume 13(4); 2015 > Article |
Abstract
Selective sweep can cause genetic differentiation across populations, which allows for the identification of possible causative regions/genes underlying important traits. The pig has experienced a long history of allele frequency changes through artificial selection in the domestication process. We obtained an average of 329,482,871 sequence reads for 24 pigs from three pig breeds: Yorkshire (n = 5), Landrace (n = 13), and Duroc (n = 6). An average read depth of 11.7 was obtained using whole-genome resequencing on an Illumina HiSeq2000 platform. In this study, cross-population extended haplotype homozygosity and cross-population composite likelihood ratio tests were implemented to detect genes experiencing positive selection for the genome-wide resequencing data generated from three commercial pig breeds. In our results, 26, 7, and 14 genes from Yorkshire, Landrace, and Duroc, respectively were detected by two kinds of statistical tests. Significant evidence for positive selection was identified on genes ST6GALNAC2 and EPHX1 in Yorkshire, PARK2 in Landrace, and BMP6, SLA-DQA1, and PRKG1 in Duroc.These genes are reportedly relevant to lactation, reproduction, meat quality, and growth traits. To understand how these single nucleotide polymorphisms (SNPs) related positive selection affect protein function, we analyzed the effect of non-synonymous SNPs. Three SNPs (rs324509622, rs80931851, and rs80937718) in the SLA-DQA1 gene were significant in the enrichment tests, indicating strong evidence for positive selection in Duroc. Our analyses identified genes under positive selection for lactation, reproduction, and meat-quality and growth traits in Yorkshire, Landrace, and Duroc, respectively.
Artificial selection is an aspect of the process of domestication development [1] that is reflected in the evolution of domestic animals. Positive selection increases the fitness of adaptive traits [2]. Previous research has demonstrated that the selection and mating of domestic animals is determined by human-generated pressures, and the genetic differences resulting from artificial selection have led to economic development [3]. Among domestic animals, the pig has experienced a particularly long history of haplotype changes through artificial selection in the process of domestication [2,4]. For most of their history, studies have found positive selection in Yorkshire, Landrace, and Duroc to be associated with specific genes related to lactation [5,6], reproduction [7], meat quality [8], and growth [3] trait. Given the importance of these traits to the pig farming industry, it is necessary to detect the signals of positive selection in pigs.
Identifying genomic selection using high-density single nucleotide polymorphism (SNP) arrays or sequencing data is useful for the discovery of putative trait-related genes. A variant under selection pressure between and/or within populations will show an increase of the frequency of particular alleles or linkage disequilibrium (LD) of SNPs. Selective sweep can cause genetic differentiation across populations [9]. The cross-population extended haplotype homozygosity (XPEHH) [10] and cross-population composite likelihood ratio (XPCLR) [9] tests are widely used to search for signals of selection between the populations. The XPEHH test detects the occurrence of selection based on measuring LD, while the XPCLR test considers the spatial patterns of allele frequencies of SNPs [9]. The XPCLR test is a likelihood method that uses differentiation of multi-locus allele frequencies between two populations to detect selective sweep. Selection with significant signals can break up the haplotype structures more rapidly than mutation or recombination processes [11].
In this study, the XPEHH and XPCLR between-population methods were implemented to detect positive selection in three pig breeds through resequencing using the Illumina HiSeq2000 platform (Illumina, Inc., San Diego, CA, USA). To further explain the biological implications of positively selected genes, functional enrichment analysis was performed.
We used genomic DNA samples gathered from 12 males and 12 females of three pig breeds: Yorkshire, 2 males and 3 females; Landrace, 7 males and 6 females; and Duroc, 3 males and 3 females. Blood samples were collected for DNA extraction. Sample collections and DNA quality check procedures were performed according to the manufacturer's instructions. Next, we constructed genomic DNA libraries for each sample using TruSeq DNA Library kits (Illumina). The paired-end library was sequenced on an Illumina HiSeq 2000 sequencing platform.
The pair-end sequence reads were aligned to the reference pig genome sequence from University of California, Santa Cruz (UCSC; http://genome.ucsc.edu/; susScr3) using Bowtie2 with the default setting. We used the following open-source software: Bowtie2, Picard tools 1.94 (http://picard.sourceforge.net), Samtools 0.1.19 [12], Genome Analysis Toolkit (GATK) 2.6.4 [13], and VCFtools 4.0 [14] for resequencing data processing and SNP calling. Substitution calling was performed using GATK Unified-Genotyper. We phased the haplotypes for the entire pig populations using BEAGLE [15]. Picard tools was used for duplicate read removal and all mate-pair information confirmation. Samtools was used for indexing the results from bam files and calculating the mapped reads using the flagstat option. GATK was used for realignment and SNP calling from resequencing data, and VCFtools was used when VCF files were handled. After filtering, non-biallelic SNPs were excluded.
To detect selective sweep, we implemented two between-populations methods (XPEHH and XPCLR) between each pairwise breed contrast. These statistics used the phased data based on the SNP genotypes obtained from the wholegenome resequencing. For each contrast, XPEHH and XPCLR were implemented across all SNPs between the three pig breeds to identify potential positive selection. We used a p-value of <0.01 for XPEHH and the top 1,000 genes in XPCLR as the cutoff criteria. These tests revealed that some SNPs might have been under positive selection in the Yorkshire, Landrace, and Duroc breeds, respectively.
Based on the findings of positive selection, enrichment analysis was performed to examine the biological functions of genes in detected regions. In this study, regions with SNPs under positive selection were extended approximately 10 kb upstream and downstream [16]. We assembled genes located within the extended region using the RefGene from the UCSC Genome Browser (http://genome.ucsc.edu/; ver. hg19). Gene enrichment analysis was performed GO terms [17], including biological process, molecular function, cellular component, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis [18] using the DAVID (http://david.abcc.ncifcrf.gov/) tool [19].
To identify non-synonymous coding SNPs (nsSNPs), we analyzed SNP effects using the SNPeffect database, which detects phenotypic effects of variation (http://snpeffect.vib.be/) [20]. For more exact results, a Fisher's exact test was performed to identify breed-specific amino acids in genes with nsSNPs. The statistical test was used to analyze 2 ├Ś 2 contingency tables composed with two factors. Considering our purpose, one factor was breed information, and the other was amino acid information. From the 2 ├Ś 2 contingency tables, we expected to observe enrichment of specific amino acids in specific breeds at each position. All data parsing and calculations were performed using Python (ver. 2.5) and R (ver. 3.1.2).
Genome-wide positive selection in pigs was identified using whole-genome resequencing, in which we aligned whole genome reads from three pig breeds, Yorkshire, Landrace, and Duroc, to the pig reference sequence. These breeds have undergone positive selection with strong signals for lactation, reproduction, meat quality, and growth. On average, ~329,482,871 sequences were mapped to the pig reference genome from the University of California, Santa Cruz (UCSC) Genome Browser (assembly ID: susScr3) using Bowtie2. Table 1 provides a summary of the alignments of 24 pig samples. Over 92.8% of all reads were aligned to the pig reference genome. Pig resequencing data with an average read depth of 11.7├Ś were generated using the Illumina HiSeq2000 platform. A total of 18,440,443 SNPs were identified using the GATK.
To detect signals of positive selection in the three pig breeds, two between-population methods (XPEHH and XPCLR) were implemented with phased haplotypes based on pairwise populations of Yorkshire-Landrace (Y-L), Yorkshire-Duroc (Y-D), Landrace-Yorkshire (L-Y), Landrace-Duroc (L-D), Duroc-Yorkshire (D-Y), and Duroc-Landrace (D-L). Here, for Y-L, Yorkshire was regarded as the observed population and Landrace as the reference population. XPEHH and XPCLR scores denote selection that happened in the observed population. As a result, several regions of positive selection were identified at a significance level of p-value < 0.01 [21] in XPEHH, and the top 1,000 genes in XPCLR. Table 2 summarizes the numbers of genes under significant positive selection, which were identified in the six breed pairs (Y-L, Y-D, L-Y, L-D, D-L, and D-Y). For example, for the Y-L breed pair, which detects selection that occurred in the Yorkshire population when compared to the Landrace population, 376 and 1,869 genes were under positive selection based on the XPEHH and XPCLR tests, respectively. For the Y-D breed pair, when using the XPCLR test, 1,498 genes were under positive selection with 3,978 SNPs detected in Yorkshire, with Duroc as the reference population. By the consensus set of genes under positive selection identified with both the XPEHH and XPCLR tests, 26, 7, and 14 genes were identified in Yorkshire, Landrace, and Duroc, respectively (Fig. 1). For example, the 26 Yorkshire genes identified by both the XPCLR and XPEHH tests in both the Y-L and Y-D comparisons, and so effectively detected four times.
We performed functional enrichment analysis on the following genes under positive selection using DAVID ver. 6.7: ST6GALNAC2 and EPHX1 in Yorkshire; PARK2 in Landrace; and BMP6, SLA-DQA1, and PRKG1 in Duroc. The results revealed enrichment of Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway terms related to metabolism, diseases, genetic or environmental information processing, and organismal systems (Table 3). The gene BMP6 acts as a hedgehog signaling pathway; the hedgehog protein is an important regulator of slow oxidative fiber clustering and is distributed throughout the tissue in pigs. Previous research has reported that BMP6 plays an important role in meat quality [8]. Among the 26 positively selected genes in Yorkshire, the ST6GALNAC2 gene is expressed during lactation, contributing to immune-related functions [5]. Up-regulation of the EPHX1 (epoxide hydrolase 1) gene occurs during mammary gland involution. Nakamura et al. [6] suggested that EPHX1 is involved in the initial stage of involution. The SEMA5A (semaphoring 5A) gene for Yorkshire reported strong candidate gene which affects the milk production traits for economic traits in cattle. The estrogen receptor (ER) gene for Landrace reported that mice lacking ER ╬▒ show severe reproduction [7]. This implies that the ER gene might be a positively selected lactation trait in pigs. The Gene Ontology (GO) terms included immune system process (SLA-DQA1) and metabolic process (PRKG1) for Duroc (Supplementary Table 2). Williams et al. [3] reported that a low level of chronic immune system resulted in greater growth in pigs. These results suggest that the 26, 7, and 14 population-specific genes with significant signals may result from potential positive selection for lactation in Yorkshire, reproduction in Landrace, and meat-quality and growth traits in Duroc.
To understand how SNPs related positive selection affect protein function, we analyzed SNP effects using the SNPeffect database. We identified two genes, SLA-DQA1 and EPHX1, with nsSNPs showing evidence of positive selection in D-Y, respectively (Fig. 2, Supplementary Table 1). To obtain accurate results, Fisher's exact test was performed on these two genes. The total numbers of statistical tests performed on the non-synonymous genes were 20 ├Ś 3 ├Ś 13 (number of cases of amino acids ├Ś number of breeds ├Ś number of non-synonymous amino acid positions) for SLA-DQA1, and 20 ├Ś 3 ├Ś 4 for EPHX1. From the enrichment tests, we identified three positions (rs324509622, rs80931851, and rs80937718) that were significant under a Bonferroni-adjusted p-value of <0.05 with an alternative hypothesis: the odds ratio is greater than 1 in the SLA-DQA1 gene. The p-values of the three non-synonymous positions were 5.20e-05, 5.20e-05, and 7.43e-06, respectively. As a note, the above three significant enrichment results were observed only in the Duroc breed. The non-synonymous positions showed strong evidence for positive selection between D-Y based on the XPEHH test. As the result, we validated the Duroc-specific enrichment of amino acids affected by non-synonymous SNPs using statistical tests. This result will contribute to improving the meat quality of pigs.
Identifying positive selection assists in the exploration of the genetic mechanisms of phenotypic diversity [2]. Selective sweep causes changes in the allele frequencies of SNPs, and the resulting changes in genetic variation can be useful in detecting selective pressure [9]. Although several studies have reported selective sweeps identified using chips such as the Illumina Porcine 60K SNP BeadChip [22] in pigs, few have identified selective sweeps using whole-genome sequencing. Therefore, detecting positive selection in the pig genome appears worthwhile.
Here we implemented two representative between-population methods, XPEHH and XPCLR, to explore positive selection. XPEHH is designed to detect near fixation or fixed selective sweeps by comparing haplotypes between two populations [10]. XPCLR is more robust for SNP discovery bias than allele frequency-based methods [9]. XPEHH tests whether the SNP site is homozygous in one population and polymorphic in another by comparing the extended haplotype homozygosity score of two populations on a core SNP. A negative XPEHH value indicates that selection happened in the reference population [15]. XPCLR was designed as a multi-locus composite likelihood ratio method to avoid the bias influence in SNP discovery. XPEHH and XPCLR find one core SNP and grid window using multi-locus information, and thus these tests identify more significant selection regions than Fst or Tajima's D [23]. Identifying significant genome-wide selection regions with multiple loci using a single-marker Fst test is difficult. Therefore, for multiple SNP methods, we carried out XPEHH and XPCLR and excluded Fst and Tajima's D. XPEHH assumes an approximately normal distribution. Park et al. [16] determined empirical p-values using the top 1% of the significance, and our results show that these significance cutoffs are more reasonable. In addition, positive selection identified by multiple test methods is more convincing.
Bioinformatic analysis of genes under positive selection in this study revealed interesting biological information related to lactation, reproduction, meat quality, and growth. We identified several GO terms using the DAVID functional annotation tool. Immune system process (GO:0002376) was found to be significant in Duroc. Williams et al. [3] reported that a low level of chronic immune system resulted in greater growth in pigs. This GO term suggests that genes with some growth traits may have been under selection in the domestication process. Growth traits are referred to as important indicators of the immune system. This suggests that SLA-DQA1 might play a role in the pig immune system. In addition, BMP6 and PRKG1 in Duroc had biological pathways related to meat-quality traits. Fonseca et al. [8] reported that the BMP6 gene plays a role in increasing the quality of meat juiciness and tenderness. Lee et al. [22] identified the significant SNPs located in the BMP6 gene related to meat quality through a genome-wide association study using the Porcine 60K BeadChip. Therefore, the SLA-DQA1 and BMP6 genes might be effects of positive selection for growth and meat quality, respectively, in Duroc. In Yorkshire, three genes under significant positive selection were ST6GALNAC2, EPHX1, and SEMA5A; these genes were related to lactation traits in several previous studies. The ST6GALNAC2 gene is expressed during lactation [5], EPHX1 is expressed in the early involution of mammary glands [6], and SEMA5A is reportedly related to milk production traits in cattle [21]. In Landrace, the ER gene was reported to be related to reproduction traits in previous studies. Generally ER refers to either ER ╬▒ or ER ╬▓ and estrogens are involved in differentiation and maintenance of reproductive tissue. Krege et al. [7] reported that mice lacking ER ╬▒ exhibited severe reproductive problems. This implies that the ER gene could be an effect of positive selection for lactation traits in pigs as well as in mice. Our findings show that several significant SNPs were present in genes found to be related to lactation, reproduction, meat quality, and growth traits in previous studies. These results indirectly support the effects of positive selection. Unfortunately, information on many of the genes with significant signals has not yet been analyzed. Based on SNP analyses for positively selected genes, SLA-DQA1 and EPHX1 genes exhibited evidence of positive selection in D-Y, respectively. However, an enrichment test revealed that the three significant amino-acid positions in SLA-DQA1 and EPHX1 were not significant.
Our analyses suggested that several genesŌĆōsuch as EPHX1, PARK2, BMP6, SLA-DQA1, PRKG1, and ST6GALNAC2ŌĆōmight have undergone positive selection. These genes are associated with lactation, reproduction, meat quality, and growth traits. All findings in this study provide useful data for the study of pigs, however our results are preliminary and need to validate these genes in another set of samples.
Acknowledgments
This work was supported by a grant from the Next-Generation BioGreen 21 Program (No. PJ008116), Rural Development Administration, Republic of Korea. All authors read and approved the final manuscript.
Supplementary materials
Supplementary data including two tables can be found with this article online at http://www.genominfo.org/src/sm/gni-13-137-s001.pdf.
Supplementary Table 1
SNP effects on population-specific genes based on the XPEHH test
Supplementary Table 2
Gene Ontology terms associated with two positively selected genes in Duroc
References
1. Hafez ES. Adaptation of Domestic Animals. Philadelphia: Lea & Febiger, 1968.
2. Larson G, Albarella U, Dobney K, Rowley-Conwy P, Schibler J, Tresset A, et al. Ancient DNA, pig domestication, and the spread of the Neolithic into Europe. Proc Natl Acad Sci USA 2007;104:15276ŌĆō15281. PMID: 17855556.



3. Williams NH, Stahly TS, Zimmerman DR. Effect of chronic immune system activation on the rate, efficiency, and composition of growth and lysine needs of pigs fed from 6 to 27 kg. J Anim Sci 1997;75:2463ŌĆō2471. PMID: 9303465.


4. Wilkinson S, Lu ZH, Megens HJ, Archibald AL, Haley C, Jackson IJ, et al. Signatures of diversifying selection in European pig breeds. PLoS Genet 2013;9:e1003453. PMID: 23637623.



5. Church PC, Goscinski A, Lefevre C. EXP-PAC: providing comparative analysis and storage of next generation gene expression data. Genomics 2012;100:8ŌĆō13. PMID: 22609187.


6. Nakamura M, Tomita A, Nakatani H, Matsuda T, Nadano D. Antioxidant and antibacterial genes are upregulated in early involution of the mouse mammary gland: sharp increase of ceruloplasmin and lactoferrin in accumulating breast milk. DNA Cell Biol 2006;25:491ŌĆō500. PMID: 16989572.


7. Krege JH, Hodgin JB, Couse JF, Enmark E, Warner M, Mahler JF, et al. Generation and reproductive phenotypes of mice lacking estrogen receptor beta. Proc Natl Acad Sci USA 1998;95:15677ŌĆō15682. PMID: 9861029.



8. Fonseca S, Wilsons IJ, Horgan GW, Maltin CA. Slow fiber cluster pattern in pig longissimus thoracis muscle: implications for myogenesis. J Anim Sci 2003;81:973ŌĆō983. PMID: 12723087.


9. Chen H, Patterson N, Reich D. Population differentiation as a test for selective sweeps. Genome Res 2010;20:393ŌĆō402. PMID: 20086244.



10. Sabeti PC, Reich DE, Higgins JM, Levine HZ, Richter DJ, Schaffner SF, et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 2002;419:832ŌĆō837. PMID: 12397357.


11. Szpiech ZA, Hernandez RD. selscan: an efficient multithreaded program to perform EHH-based scans for positive selection. Mol Biol Evol 2014;31:2824ŌĆō2827. PMID: 25015648.



12. Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009;25:2078ŌĆō2079. PMID: 19505943.



13. McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010;20:1297ŌĆō1303. PMID: 20644199.



14. Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, et al. The variant call format and VCFtools. Bioinformatics 2011;27:2156ŌĆō2158. PMID: 21653522.



15. Browning BL, Browning SR. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am J Hum Genet 2009;84:210ŌĆō223. PMID: 19200528.



16. Park W, Kim J, Kim HJ, Choi J, Park JW, Cho HW, et al. Investigation of de novo unique differentially expressed genes related to evolution in exercise response during domestication in Thoroughbred race horses. PLoS One 2014;9:e91418. PMID: 24658125.



17. Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R, et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res 2004;32:D258ŌĆōD261. PMID: 14681407.



18. Kanehisa M, Goto S, Kawashima S, Nakaya A. The KEGG databases at GenomeNet. Nucleic Acids Res 2002;30:42ŌĆō46. PMID: 11752249.



19. Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, et al. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol 2003;4:P3. PMID: 12734009.


20. Reumers J, Schymkowitz J, Ferkinghoff-Borg J, Stricher F, Serrano L, Rousseau F. SNPeffect: a database mapping molecular phenotypic effects of human non-synonymous coding SNPs. Nucleic Acids Res 2005;33:D527ŌĆōD532. PMID: 15608254.


21. Pareek CS. Emerging trends on functional genomics in farm animals. (Parihar P, ed.). In: Advances in Applied Biotechnology Jodhpur: Agrobios, 2010. pp. 23ŌĆō34.
22. Lee T, Shin DH, Cho S, Kang HS, Kim SH, Lee HK, et al. Genome-wide association study of integrated meat quality-related traits of the duroc pig breed. Asian-Australas J Anim Sci 2014;27:303ŌĆō309. PMID: 25049955.




23. Ma Y, Zhang H, Zhang Q, Ding X. Identification of selection footprints on the X chromosome in pig. PLoS One 2014;9:e94911. PMID: 24740293.



Fig.┬Ā1
Visualization of genome-wide positive selection. Manhattan plots of the degree of selection signals based on pairwise populations of three pig breeds using cross-population extended haplotype homozygosity (A) and cross-population composite likelihood ratio (B). Red lines indicate the significance level and colored dots indicate the genes under significant positive selection. Y-L, Yorkshire-Landrace; Y-D, Yorkshire-Duroc; L-D, Landrace-Duroc; L-Y, Landrace-Yorkshire; D-L, Duroc-Landrace; D-Y, Duroc-Yorkshire.

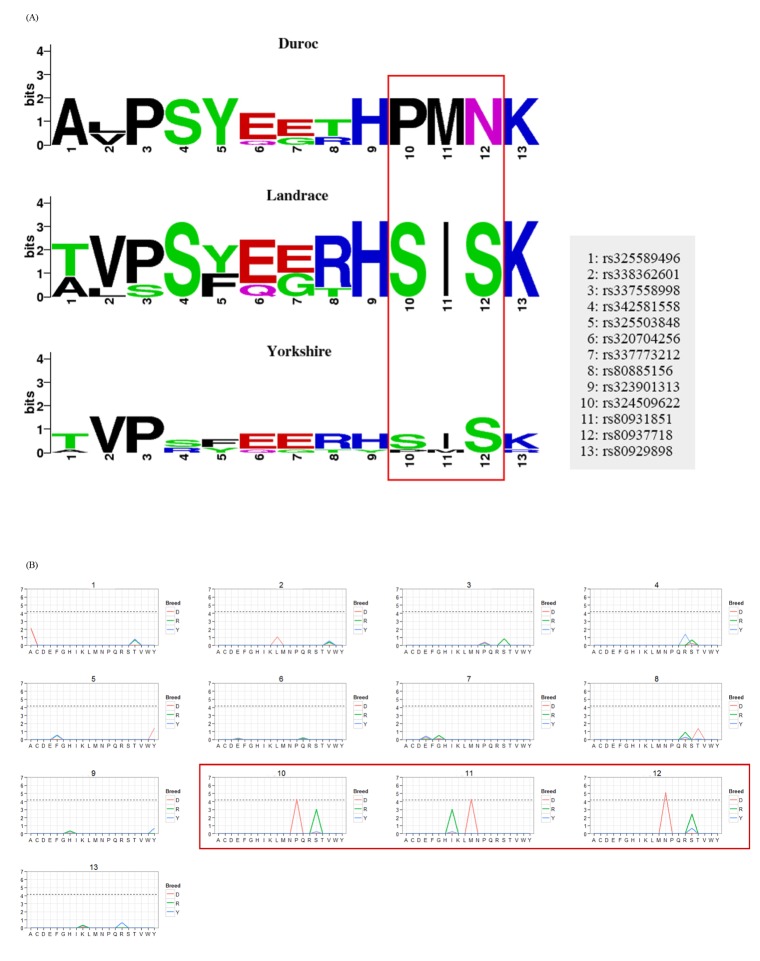
Fig.┬Ā2
Visualization of single nucleotide polymorphism (SNP) effects on positively selected genes. (A) Visualization of the sequence logo for the SLA-DQA1 gene with 13 non-synonymous SNPs (nsSNPs) showing evidence of positive selection in Duroc. (B) Visualization of enrichment tests for the 13 nsSNPs. The red box indicates the significant positive selection regions under the Bonferroni adjusted p-value < 0.05.
Table┬Ā1.
Summary of alignment rates in 24 pig samples
Table┬Ā2.
Summary of genes under significant positive selection
| Breed pair | XPEHH | XPCLR | Overlapping genes | Gene names for breed specific genes | |
|---|---|---|---|---|---|
| Yorkshire (Y) | Y_L | 376 (26,458) | 1,869 (4,848) | 119 | 26 (ST6GALNAC2a, TBC1D13, ARHGAP26, TRAPPC12, RPS7, COLEC11, DEPTOR, ENPP2, OLFM3, EXTL2, CLEC2D, DUS2, CHST9, HOOK1, PDLIM5, TMEM63A, EPHX1a, MGAT5B, OSBPL10, COL13A1, ADAMTS14, TBATA, NUP155, SEMA5Aa, PHACTR3, LRGUK) |
| Y_D | 336 (24,887) | 1,498 (3,978) | 99 | ||
| Landrace (L) | L_Y | 756 (47,167) | 1,739 (4,751) | 69 | 7 (MAS1, ZC3H12D, EEA1, GLG1, ERa, PARK2a, IDO1) |
| L_D | 408 (26,698) | 1,733 (4,539) | 143 | ||
| Duroc (D) | D_L | 495 (76,009) | 2,108 (5,327) | 60 | 14 (TGFBI, LMTK2, PRDM14, BMP6a, SLA-DQB2, SLADQA1a, DST, TBC1D14, CWH43, ENOPH1, GOLT1A, PRKG1a, LIPM, ANKRD22) |
| D_Y | 587 (68,590) | 2,032 (5,364) | 64 |
The single nucleotide polymorphisms under positive selection were extended by approximately ┬▒10 kb.
Parentheses indicate the number of single nucleotide polymorphisms at the significance level.
XPEHH, cross-population extended haplotype homozygosity; XPCLR, cross-population composite likelihood ratio.
Table┬Ā3.
Enriched KEGG pathways associated with positively selected genes
| Breed | Gene symbol | KEGG pathway | Higher-level KEGG pathway | The highest level of KEGG pathway |
|---|---|---|---|---|
| Yorkshire | ST6GALNAC2 | Glycosphingolipid biosynthesis | Glycan biosynthesis and metabolism | Metabolism |
| EPHX1 | Metabolism of xenobiotics by cytochrome P450 | Xenobiotics biodegradation and metabolism | ||
| Landrace | PARK2 | Parkinson's disease, | Neurodegenerative diseases | Human diseases |
| Ubiquitin mediated proteolysis | Folding, sorting and degradation | Genetic information processing | ||
| Duroc | BMP6 | Hedgehog signaling pathway [8, 22] | Signal transduction | Environmental information processing |
| TGF-beta signaling pathway | ||||
| SLA-DQA1 | Cell adhesion molecules | Signaling molecules and interaction | Environmental information processing | |
| Antigen processing and presentation, intestinal immune network for IgA production | Immune system | Organismal systems | ||
| Type I diabetes mellitus | Endocrine and metabolic diseases | Human diseases | ||
| Asthma, autoimmune thyroid disease, systemic lupus erythematosus, allograft rejection, graft-versus-host disease | Immune diseases | Immune diseases | ||
| Viral myocarditis | Cardiovascular diseases | Human diseases | ||
| PRKG1 | Long-term depression | Nervous system | Organismal systems | |
| Olfactory transduction | Sensory system | |||
| Gap junction | Cell communication | Cellular processes |
- TOOLS
-
METRICS

-
- 4 Crossref
- 6,594 View
- 80 Download





