 |
 |
- Search
| Genomics Inform > Volume 18(4); 2020 > Article |
|
Abstract
The coronavirus disease 2019 is a contagious disease and had caused havoc throughout the world by creating widespread mortality and morbidity. The unavailability of vaccines and proper antiviral drugs encourages the researchers to identify potential antiviral drugs to be used against the virus. The presence of RNA binding domain in the nucleocapsid (N) protein of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) could be a potential drug target, which serves multiple critical functions during the viral life cycle, especially the viral replication. Since vaccine development might take some time, the identification of a drug compound targeting viral replication might offer a solution for treatment. The study analyzed the phylogenetic relationship of N protein sequence divergence with other 49 coronavirus species and also identified the conserved regions according to protein families through conserved domain search. Good structural binding affinities of a few natural and/or synthetic phytocompounds or drugs against N protein were determined using the molecular docking approaches. The analyzed compounds presented the higher numbers of hydrogen bonds of selected chemicals supporting the drug-ability of these compounds. Among them, the established antiviral drug glycyrrhizic acid and the phytochemical theaflavin can be considered as possible drug compounds against target N protein of SARS-CoV-2 as they showed lower binding affinities. The findings of this study might lead to the development of a drug for the SARS-CoV-2 mediated disease and offer solution to treatment of SARS-CoV-2 infection.
The outbreak of novel coronavirus infection has drastically affected the lives of the human population worldwide. This infection started as respiratory illness/pneumonia of unknown origin in Wuhan city of China at the end of the year 2019. The organism identified and termed as novel on 7 January 2020. The World Health Organization (WHO) declared it as a public health emergency of international concern as the disease spread to other regions of the world [1]. The official name of this infection was made as coronavirus disease 2019 (COVID-19) on 11 February 2020. The epidemic was declared a pandemic officially by WHO on 11 March 2020. The novel coronavirus is also termed, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [1]. SARS-CoV-2 infection mainly causes pneumonia, upper and lower respiratory tract infection with fever and cough as significant clinical symptoms. But some other symptoms include shortness of breath, muscle pain, confusion, headache, sore throat, and acute respiratory distress syndrome, leading to respiratory or multi-organ failure including renal and neurological diseases [2,3].
Coronaviruses (CoVs) are a group of large enveloped viruses with positive sense, single-stranded RNA genomes. Previously identified CoVs in human disease are the alpha CoVs (hCoV-NL63, hCoV-229E) and the beta CoVs (hCoV-0C43), severe acute respiratory syndrome CoV (SARS-CoV), and the Middle East respiratory syndrome CoV (MERS-CoV) [4]. However, among these emerging, highly pathogenic human CoVs, SARS-CoV, MERS-CoV and the newly emerged SARS-CoV-2 infection can result in life-threatening disease conditions and the potential to cause pandemic [2].
The outcome of SARS-CoV-2 sequencing (NCBI reference sequence: NC_045512.2) has proposed about the significant sequence level identity of SARS-CoV-2 with SARS-CoV (79%) rather than MERS-CoV (50%). Besides, the higher levels of transmissibility and pandemic risk of COVID-19 at an early stage has been reported in many studies [1]. In the available literatures, the size of the SARS-CoV-2 (NCBI reference sequence: NC_045512.2) genome is 30KB. The genomic virion consists of four major protein regions including matrix (M) protein, an envelope (E) protein, spike (S) protein, and a nucleocapsid (N) protein within the viral envelope [5,6]. The functional architectures of each of these viral proteins have accurately characterized. S protein primarily binds to the host cell receptor and form attachment with the host body. Alternatively, M and E proteins are involved in the formation of the viral envelope [6]. Similarly, SARS-CoV-2 protein N is a multifunctional RNA binding protein, necessary for viral RNA transcription, replication and/or assembly of virus [6]. Interestingly, a unique N-terminal RNA binding domain of SARS-CoV-2 N protein has identified as a novel antiviral drug target site [7]. The viral N protein packages the genome into long, flexible, and helical RNP complexes, called nucleocapsids which protect the SARS-CoV-2 virion structure [5]. Additionally, N protein has a significant contribution towards timely replication and reliable transmission of SARS-CoV-2 during its life cycle. Therefore, N protein (PDB ID: 6VYO) can be considered as a novel drug target of SARS-CoV-2.
The SARS-CoV-2 infection has created a dangerous pandemic situation due to its quick transmission and deadly nature. It has affected both the health and economy of human population across the globe tremendously. Many ongoing pieces of research are trying to develop vaccines to control this situation, but all are in various phases of trials. Thus, the present study has focused on in silico discovery of potent leads from several antiviral drugs and compounds of plant origin against SARS-CoV-2 infection. The present study would throw lights on the discovery of antiviral drug against SARS-CoV-2.
Nucleocapsid protein sequences of total 49 CoV species and/or strains including SARS-CoV-2 were retrieved in FASTA format from NCBI web server (https://www.ncbi.nlm.nih.gov/) on 30 March 2020. Two N proteins of Ebola and H1N1 virus were included to study evolutionary divergence across species. Further, total 51 N protein sequences were aligned using MUSCLE algorithm of Molecular Evolutionary Genetics Analysis 7 (MEGA 7) package [8]. The resulted alignment was used to generate phylogenetic tree using neighbour joining (NJ) method of MEGA 7 for 1,000 bootstrap replicates.
Functional domains of SARS-CoV-2 N protein (YP_009724397.2) were identified using NCBI conserved domain database (CDD) (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) search. The CDD is a collection of domain models which imports information from Pfam, SMART, COG, and NCBI to provide a more accurate assessment of neighbor relationships between protein sequences [9].
The secondary structure of SARS-CoV-2 N protein was predicted from its complete amino acid sequence (accession No. YP_009724397.2) using PSIPRED 4.0 algorithm [10]. Similarly, protein disorder portion and membrane helix region was predicted by using DISOPRED3 [10] and MEMSAT-SVM algorithm [10] of PSIPRED web server (http://bioinf.cs.ucl.ac.uk/psipred/).
Available N-terminal domain structure (PDB ID: 6VYO) of SARS-CoV-2 N protein was retrieved from Protein Data Bank (PDB) (https://www.rcsb.org/). Initially, hydrogen atoms were added to protein structure after removal of all water and other hetero molecules. Further, energy minimization was performed using Discovery Studio 3.5 suite to obtain a properly optimized structure of target protein.
In absence of knowledge on exact drug-binding site, probable binding cavity within SARS-CoV-2 N protein was predicted using metaPocket 2.0 (https://projects.biotec.tu-dresden.de/metapocket/). MetaPocket tool identifies cavities on protein surface for drug-binding site prediction using multiple computational approaches [11] such as PASS11, LIGSITE, Fpocket, SURFNET, GHECOM, and ConCavity.
Different natural compounds of plant origin reported with antiviral, anti-inflammation, anti-influenza, antiŌĆÆhuman immunodeficiency virus, anti-hepatic properties were shortlisted from different literatures. In addition, few Food and Drug Administration approved, and investigational antiviral drugs were also selected from Drug Bank (https://www.drugbank.ca/) database for further investigation.
Three-dimensional structures of natural ligands were retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/) database in SDF format and converted into PDB format using Discovery Studio 3.5 suite. Similarly, PDB structures of antiviral drugs were collected from the Drug Bank (https://www.drugbank.ca/). Further, structure optimization and protonation state of all ligands were achieved using Discovery Studio 3.5 suite.
Molecular docking was performed between all selected ligands (phytochemicals and antiviral drugs) and the drug target (N protein, PDB ID: 6VYO) separately in order to identify the most efficient inhibitor against SARS-CoV-2. AutoDock 4.2 (http://autodock.scripps.edu/) and AutoDock Tools 4 tool [12] were used to perform molecular docking study. The N-terminal RNA binding domain of SARS-CoV-2 N protein was observed as a homotetramer structure; therefore, only chain A of the available crystal structure was employed for docking analysis. Prior to docking, Kollman charges and polar hydrogen atoms were added to the target structure. Both ligand and receptor structures were prepared using ADT tool and converted to pdbqt format before docking. A virtual grid box was set around the drug-binding cavity of the target structure with size of 74, 78, and 74 ├ģ in x, y, and z direction in spacing of 0.375 ├ģ. Semi flexible docking was performed by maintaining target structure as rigid and allowing flexibility to ligand molecules within the drug-binding pocket [13]. Lamarckian genetic algorithm was used with 25,000,000 energy evaluation steps for each dock run. Auto dock generated 10 conformers based on free binding energy for each protein-ligand complex. The most energetically favorable (lowest energy) binding complex was considered for analysis. Further analysis and presentation of atomic interaction between docked complexes were performed using PyMol molecular graphics tool (http://www.pymol.org).
Total 49 N proteins different CoV species, including SARS-CoV-2 (Table 1) were retrieved to construct the phylogenetic tree. Again, protein sequences of two distance homologues of SARS-CoV-2 such as Ebola (accession No. SCD11531.1) and H1N1 (accession No. YP_009118629.1) virus were included within the tree in order to establish sequential divergence pattern across species. The phylogenetic tree was constructed using NJ method [14] with tree evaluation step for 1,000 bootstrap replicates. The resulted rooted tree (Fig. 1) clustered into two major clades. Total 49 species were diversified within both of the clades (clade-I, 26; clade-II, 23). The target N protein sequence of SARS-CoV-2 (accession No. YP_009724397.2) was grouped with SARS-CoV (severe acute respiratory syndrome-related virus) (accession No. NP_828858.1) sequence within clade-I with branch frequency of 100% which pointed out regarding their significant evolutionary closeness. One separate clade was formed within the tree with branch frequency of 61% among the two outgroups (Ebola and H1N1) which clearly revealed their divergence from all other 49 sequences.
The complete sequence of SARS-CoV-2 N protein (accession No. YP_009724397.2) comprises of 419 amino acids. All functional domain regions within the N protein sequence of SARS-CoV-2 were identified from its conserved pattern among the members of beta CoV nucleocapsid protein family. The conserved domains were observed within the aligned region of SARS-CoV-2 N protein from 14ŌĆÆ368 amino acids (Fig. 2A) with the members of the superfamily (pfam00937) (Fig. 2B). The CD search identified one. N-terminal (50ŌĆÆ175 amino acids) and one C-terminal (258ŌĆÆ359 amino acids) functional domain (Fig. 2C) with good bit score (424.07) and lowest e-value (7.05e-148). The nucleocapsid N-terminal domain (NTD) of SARS-CoV-2 was showed significant similarities with the conserved domain of family cd21554 whereas the C-terminal domain (CTD) found conserved within the family members of cd21595 (Fig. 2D).
In the absence of full-length structure, the secondary structural elements of SARS-CoV-2 N protein were predicted from its primary sequence using PSIPRED web server. Secondary structural elements such as two long, eight medium, two short helical regions and two medium, nine short ╬▓-sheets were predicted within the complete sequence of SARS-CoV-2 N protein (Fig. 3).
Most of the NTD (50ŌĆÆ175) regions were predicted as ╬▓-sheets and coils. On the contrary, structural elements such as helices, ╬▓-sheets, and coils were observed within CTD (258ŌĆÆ359) regions (Fig. 3). Further, highly disordered regions of SARS-CoV-2 N protein were observed above the cut off score (0.5) from amino acid positions 1ŌĆÆ50, 180ŌĆÆ250, and 350ŌĆÆ419 (Fig. 4A). However, significant disorder portions were absent within the both NTD (50ŌĆÆ175) and CTD (258ŌĆÆ359) regions (Fig. 4A). According to MEMSAT-SVM algorithm, the sub-cellular localization of SARS-CoV-2 nucleocapsid NTD was found as cytoplasmic, whereas a small C-terminal transmembrane region was noticed from 302ŌĆÆ317 amino acids (Fig. 4B).
Homology search using BLASTP algorithm revealed the structure of N-terminal RNA binding domain occupied 30% region of SARS-CoV-2 N protein (accession No. YP_009724397.2) sequence with 100% identity. Therefore, the three-dimensional structure of SARS-CoV-2 N protein was retrieved and processed for structural correction and optimization. The possible drug-binding cavity of SARS-CoV-2 N protein was predicted in the absence of literary evidence. Algorithm of metaPocket was generated top three hits after clustering the results of PASS11, LIGSITE, Fpocket, SURFNET, GHECOM, and ConCavity. Out of these three, the large active pocket was considered a possible drug-binding cavity (Fig. 5).
As of literature, a total of eight natural compounds of plant origin and three synthetic compounds (Table 2) were identified with antiviral properties, therefore, prepared to dock against SARS-CoV-2 N protein.
Again, seven antiviral drugs (Table 3) were also included within the study to discover potent inhibitor against N protein of SARS-CoV-2. Finally, 3D structures of a total of eighteen ligands were extracted from online databases (PubChem/Drug Bank) and prepared for docking study.
Molecular docking is an efficient technique to identify the binding affinity of a drug compound against a drug target [15,25]. Therefore, all possible inhibitors were docked separately against SARS-CoV-2 N protein to discover effective ligand and important atomic interaction between protein-ligand complexes within the drug-binding cavity. The resulted in free binding energy, and the inhibition constant of each binding complex was reported in Table 4. According to docking energy score and inhibition constant (KI), total eight antiviral compounds such as glycyrrhizic acid (ŌĆÆ12.61 kcal/mol; KI, 573.72 pm), theaflavin (ŌĆÆ10.35 kcal/mol; KI, 26.03 nM), diosgenin (ŌĆÆ10.06 kcal/mol; KI, 42.53 nM), U18666A (ŌĆÆ9.08 kcal/mol; KI, 219.38 nM), ethyl brevifolincarboxylate (ŌĆÆ9.07 kcal/mol; KI, 226.42 nM), quercitrin (ŌĆÆ9.04 kcal/mol; KI, 238.18 nM), curcumin (ŌĆÆ8.68 kcal/mol; KI, 434.59 nM), and ladanein (ŌĆÆ8.19 kcal/mol; KI, 988.63 nM) showed good binding efficiency than rest of the compounds (Table 4). Presence of an ample number of polar interactions has a significant contribution towards the stability of a specific ligand within the binding site of drug target. Therefore, h-bond interaction between the drug target and ligands were inspected. Interestingly, good binding affinity and strong h-bond interaction within distance Ōēż 3.5 ├ģ from binding cavity were identified in case of 10 suitable compounds such as glycyrrhizic acid (ŌĆÆ12.61 kcal/mol; h-bond, 16 nos), theaflavin (ŌĆÆ10.35 kcal/mol; h-bond, 11 nos), ethyl brevifolincarboxylate (ŌĆÆ9.07 kcal/mol; h-bond, 6 nos), quercitrin (ŌĆÆ9.04 kcal/mol; h-bond, 11 nos), curcumin (ŌĆÆ8.68 kcal/mol; h-bond, 5 nos), ladanein (ŌĆÆ8.19 kcal/mol; h-bond, 8 nos), apigenin (ŌĆÆ7.98 kcal/mol; h-bond, 6 nos), tenofovir (ŌĆÆ6.92 kcal/mol; h-bond, 9 nos), resveratrol (ŌĆÆ6.91 kcal/mol; h-bond, 5 nos), ribavirin (ŌĆÆ6.41 kcal/mol; h-bond, 12 nos), indicated about their efficacy to block the important site within the RNA binding domain of SARS-CoV-2 N protein (Tables 4 and 5, Fig. 6).
To its support, few amino acid residues such as PHE 66, PRO 67, ARG 68, GLY 69, GLN 70, TYR 123, TRP 132, and ALA 134 were found commonly interacting with all of these ligands within the binding cavity of SARS-CoV-2 N protein. However, presence of h-bond interaction with quite good binding energy and inhibition constant values were also noticed in case of rest seven antiviral compounds such as diosgenin (ŌĆÆ10.06 kcal/mol; KI, 42.53 nM; h-bond, 3 nos), U18666A (ŌĆÆ9.08 kcal/mol; KI, 219.38 nM; h-bond, 2 nos), berberine (ŌĆÆ7.87 kcal/mol; KI, 1.69 ┬ĄM; h-bond, 2 nos), emodin (ŌĆÆ7.82 kcal/mol; KI, 1.86 uM; h-bond, 6 nos), quercetin (ŌĆÆ7.47 kcal/mol; KI, 3.33 ┬ĄM; h-bond, 8 nos), hydroxychloroquine (ŌĆÆ7.35 kcal/mol; KI, 4.07 ┬ĄM; h-bond, 2 nos), chloroquine (ŌĆÆ6.86 kcal/mol; KI, 9.34 ┬ĄM; h-bond, 1 nos) inbound form with SARS-CoV-2 N protein (Table 4, Fig. 7). Overall docking study confirmed the binding potential of the discussed phytochemicals and drugs, against drug target, Nucleocapsid protein of SARS-CoV-2.
The SARS-CoV-2 or COVID-19 pandemic has created an alarming situation due to severe infection and death rate worldwide. Researchers all over the world are in search to identify novel drug/vaccine target as well as the development of drug/vaccine to combat the disease. Several recent studies have been reported probable synthetic drug candidates such as conivaptan, amyrin, ZINC000027115482 [26], ritonavir, lopinavir, umifenovir [27], theophylline, pyrimidine [28], simeprevir and grazoprevir [29] against nucleocapsid protein of SARS-CoV-2. As, N protein has a vital role for the survival and growth of SARS-CoV-2 thus authors focused on the discovery of potential natural or synthetic compounds to block its regular mechanism. In support of the present scenario, the current study has tried to conduct some critical analyses on important drug target, i.e., nucleocapsid (N) protein of SARS-CoV-2. The present research also focuses on in silico discovery of potent natural/synthetic compounds against the virus.
The phylogenetic study among different CoV species community identified the close relation and less diversification between N proteins of SARS-CoV and SARS-CoV-2, which indicates the high similarities between those species. The protein family sequence similarity search or the conserved domain search points out the versatility of SARS-CoV-2 N protein, which is predicted by the conserved amino acid regions from different members CoV superfamilies such as SARS-CoV, murine CoV (murine hepatitis virus) and alpha CoV-1 species (Feline infectious peritonitis virus).
Primary sequence analysis resulted in two crucial functional domain regions both in N and C terminals of SARS-CoV-2. Interestingly, the NTD comprises RNA binding site, which signifies its importance towards a viral cellular mechanism. To its support, the available crystal structure of NTD SARS-CoV-2 N protein was retrieved and utilized in further study. The SARS-CoV-2 N protein had no binding site information including drug-binding sites till the end of March 2020, which influences the researchers to predict the drug-binding pocket in RNA binding domain of N protein. But recently, Kang et al. [30] reported about the crystal structure and showed the drug-binding pocket (including the amino acids Tyr 110, Tyr 112, Tyr 55, Ala56, and Arg89) of N protein with PDB ID 6M3M whereas this present study predicted the binding domain in SARS-CoV-2 N protein (PDB ID: 6VYO) with amino acids positioned from 64ŌĆÆ71, 84, 123ŌĆÆ124, and 131ŌĆÆ140. This study represents the maximum similarities between the crystal structure binding pocket and the presently identified drug-binding pocket in N protein, which should be considered while deciding a drug for trial in the treatment of the disease.
Today, the death report of COVID-19 from different corner of the globe is drastically increasing due to the absence of an effective antiviral drug. To overcome this situation, eighteen compounds, including natural compounds of plant origin and antiviral drugs, were docked into the drug-binding cavity of N protein to identify potential ligands against SARS-CoV-2. This study has been able to find the binding efficiency of a few phytochemicals (Theaflavin, curcumin, ladanein), and a few drug compounds (glycyrrhizic acid, ethyl brevifolin caboxylate, and quercitrin) against N protein of the virus. This might serve as information about their potential to be a treatment option for SARS-CoV-2. The antiviral effects of phytochemicals such as Theaflavin, curcumin, and ladanein, against many pathogenic viruses, have already been well studied and reported. Theaflavin is known to prevent from influenza virus by inhibiting its replication [15].
Similarly, curcumin has antiviral properties against H1N1 Influenza and FIPV [16]. Again, the inhibitory effect of ladanein against hepatitis C virus infection [17] is also well studied. Thus, these compounds may be useful as an anti-infective agent against COVID-19. Antiviral drugs such as glycyrrhizic acid, ethyl brevifolincarboxylate, and quercitrin have inhibitory effect against [18,23] hepatitis B and C virus. But, glycyrrhizic acid and quercetin are associated with severe side effects such as hypokalemia, oedema, rhabdomyolysis or myoglobinuria, mitochondrial toxicity, and mutagenicity [31,32]. However, according to the resulted binding affinities and the presence of H-bonds glycyrrhizic acid and theaflavin can be considered as suitable drug compounds against SARS-CoV-2 N protein. In regards to toxicity associated with glycyrrhizic acid, the use of natural compound, i.e., theaflavin may be more effective against COVID-19. Other than the mentioned natural/synthetic compounds, few others such as diosgenin [17], U18666A [19], apigenin (Ocimum sanctum) [20], resveratrol (Vitis labrusca) [21], berberine (Berberis vulgaris) [22], emodin (Radix et Rhizoma Rhei, Radix Polygoni Multiflori) [24], and tenofovir (Phyllanthus niruri) [18] has shown stable binding interaction with SARS-CoV-2 N protein. Hence they may also be studied for further validation.
The COVID-19 outbreak has caused havoc throughout the world, changing the course of human lives. Researchers are trying to design a vaccine against SARS-CoV-2 but that might take some time. This study attempts to find a drug for treating the disease condition, which will help to save human lives and mitigate the sufferings of millions of people infected by the virus worldwide. Some antivirals phytocompounds and synthetic drugs have been analyzed in this in silico study, which would target the N protein, responsible for replication of SARS-CoV-2 in the host body. Of all the compounds in this study, glycyrrhizic acid and theaflavin can be used as the antiviral drug, as they showed a higher binding affinity with the target protein. The effective drug candidates would be helpful to prevent the SARS-CoV-2 viral N protein and to reduce the risk of infection in the host body.
Notes
Acknowledgments
We are thankful to Dr. Pawan Kumar Agrawal, Vice chancellor, Odisha University of Agriculture and Technology for his moral support and valuable suggestions.
Fig.┬Ā1.
Phylogenetic tree were presented among 49 nucleocapsid (N) protein sequences of severe acute respiratory syndrome coronavirus (SARS-CoV) and SARS-CoV-2 from different species. The number in the left side of tree denotes bootstrap frequency for each taxon. The N protein of out group (Ebola and H1N1) sequences and the target SARS-CoV-2 protein were highlighted using red and blue outline respectively. Similarly, N protein sequence of SARS-CoV was highlighted using green outline.
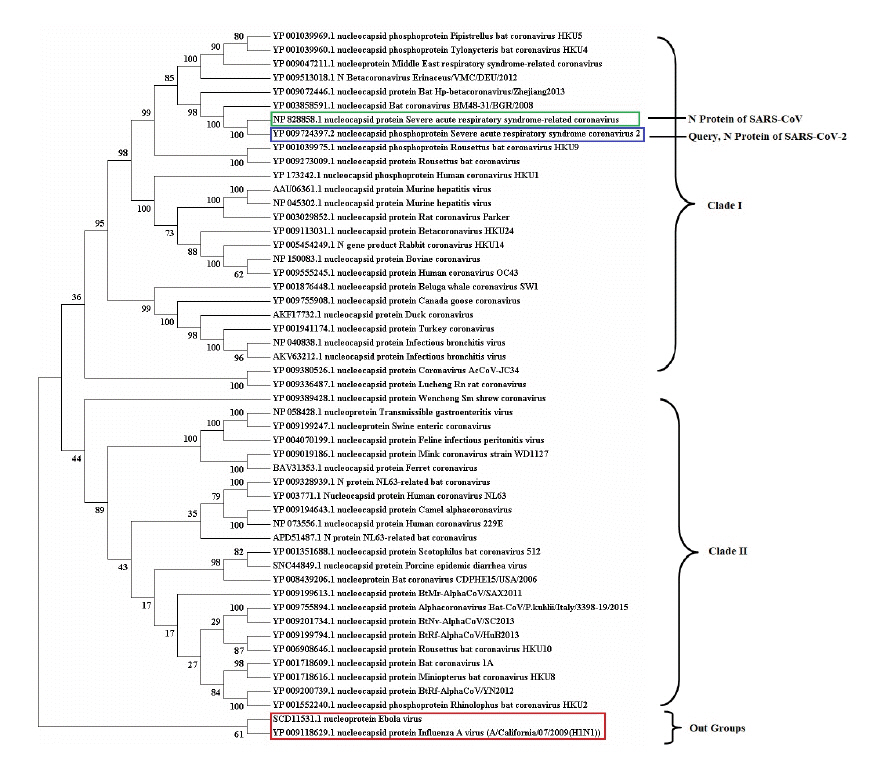
Fig.┬Ā2.
Conserved functional domains of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid protein. (A) Sequence alignment between SARS-CoV-2 and members of super family (pfam00937). (B) The alignment between SARS-CoV-2 and consensus sequence of pfam00937 nucleocapsid protein. The conserved amino acid patterns were highlighted using boxes. (C) All functional domain regions of SARS-CoV-2 nucleocapsid protein were presented in schematic diagram. N-NTD, nucleocapsid protein N-terminal domain; N-CTD, nucleocapsid protein C-terminal domain. (D) The sequence alignment of N-NTD (50-175) and N-CTD of SARS-CoV-2 with their respective conserved domain family.
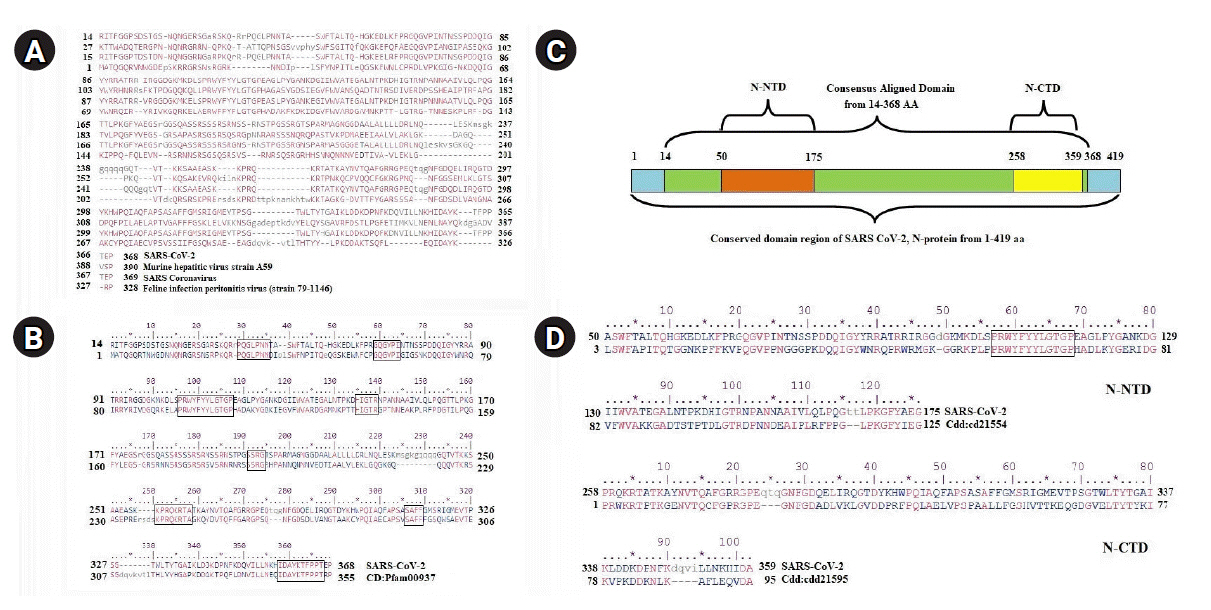
Fig.┬Ā3.
Predicted secondary structural elements for full length nucleocapsid protein of severe acute respiratory syndrome coronavirus 2. Helix, pink cylinder; Sheet, yellow cylinder.

Fig.┬Ā4.
(A) The disorder plot of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid protein was deciphered. X-axis: amino acid residue number; Y-axis: disorder cut off value. Black color dots were used to plot disorder values on the Y-axis for the corresponding amino acids on X-axis. (B) Representation of sub-cellular localization of SARS-CoV-2 nucleocapsid N-terminal domain.
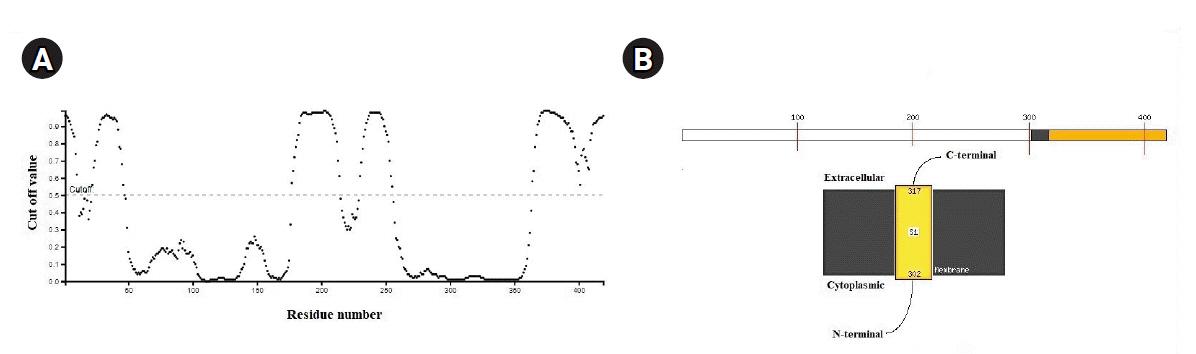
Fig.┬Ā5.
(A) Cartoon representation of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid protein (PDB ID: 6VYO, chain A) structure. Β-sheet, pink colour arrows; Coil, tube. (B) Space filling representation. Active drug-binding pocket was highlighted using red colour within the structure.
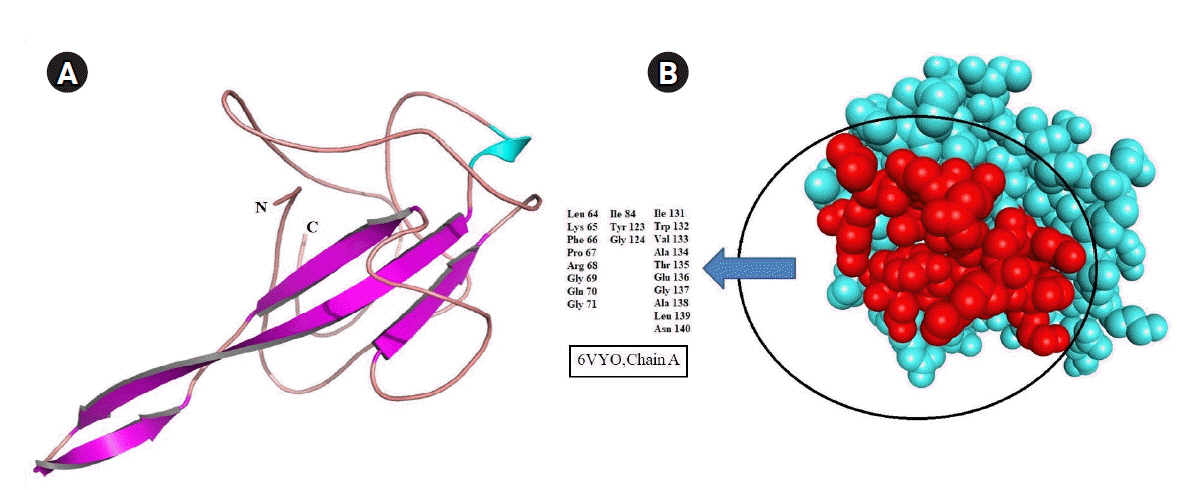
Fig.┬Ā6.
Polar interaction between severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid protein with natural/synthetic compounds: glycyrrhizin (A), theaflavin (B), ethylbrevifolincarboxylate (C), quercitrin (D), curcumin (E), ladanein (F), apigenin (G), tenofovin (H), resveratrol (I), and ribavirin (J).
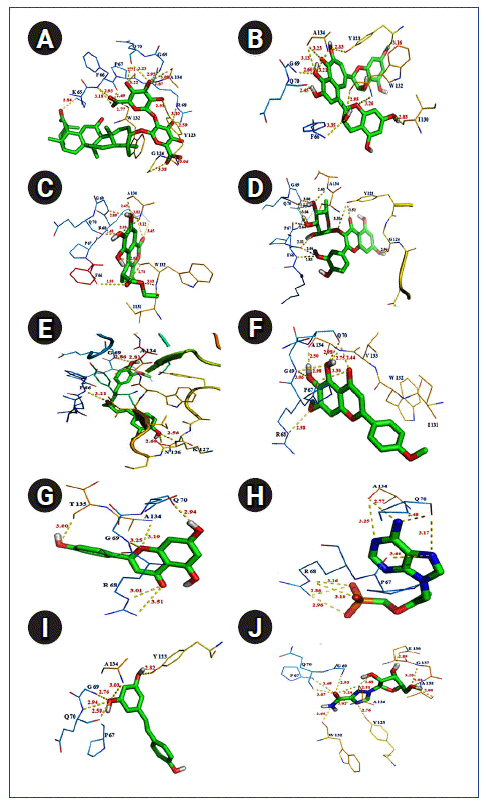
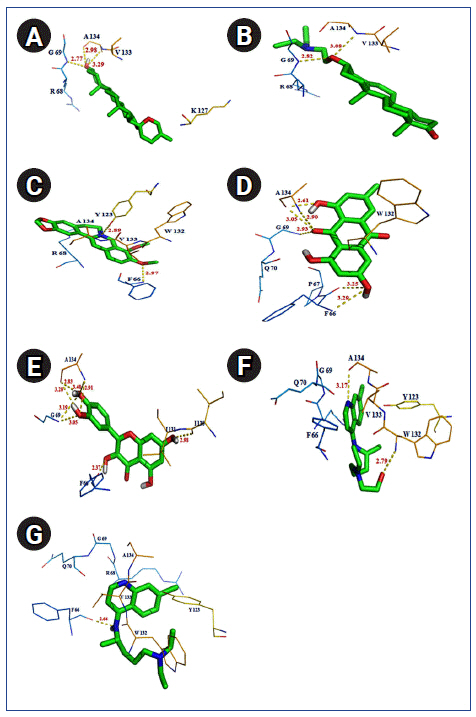
Fig.┬Ā7.
Polar interaction between severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) nucleocapsid protein with natural/synthetic compounds: diosgenin (A), U18666A (B), berberine (C), emodin (D), quercetin (E), hydroxyl chloroquine (F), and chloroquine (G).
Table┬Ā1.
Nucleocapsid proteins from different coronavirus species collected from NCBI
Table┬Ā2.
Eleven ligand molecules (natural and synthetic) and antiviral properties
| No. | Natural/Synthetic compounds | Pub Chem CID | Source/Plant name | Property | Virus | Reference |
|---|---|---|---|---|---|---|
| 1 | Theaflavin | 135403798 | Camellia sinensis (tea plant) | Prevents influenza by inhibiting replication using potentially directs virucidal effect | H1N1 | [15] |
| 2 | Curcumin | 969516 | Curcuma longa L. (turmeric) | Antiviral activity against FIPV | FIPV, HIV, influenza | [16] |
| Inhibition of HIV-1 and HIV-2 proteases | ||||||
| Inhibition of haemagglutination | ||||||
| 3 | Diosgenin | 99474 | Synthetic | Effectively blocks the replication of hepatitis C virus | Hepatitis C virus | [17] |
| 4 | Ladanein | 3084066 | Marrubium peregrinum L. | Effectively inhibits the post attachment entry step of hepatitis C virus | Hepatitis C virus | [17] |
| 5 | Quercetin | 5280343 | Phyllanthus niruri | Inhibits virus replication and viral nucleocapsid formation by inhibiting DNA polymerase of hepatitis B | Hepatitis B/C virus | [17,18] |
| 6 | Ethyl brevifolincarboxylate | 5487248 | Synthetic | Inhibits virus replication and viral nucleocapsid formation by inhibiting DNA polymerase of hepatitis B | Hepatitis B virus | [18] |
| 7 | Quercitrin | 5280459 | Phyllanthus niruri | Inhibit virus replication and viral nucleocapsid formation by inhibiting DNA polymerase of hepatitis B | Hepatitis B virus | [18] |
| 8 | U18666A | 9954082 | Synthetic | Inhibits the proliferation of type 1 FIPV | Feline infectious peritonitis virus | [19] |
| 9 | Apigenin | 5280443 | Ocimum sanctum (Tulsi) | Prevents the early multiplication of H1N1 virus and control the viral growth | H1N1 | [20] |
| 10 | Resveratrol | 445154 | Vitis labrusca | Effectively reduce the inflammatory cell production and pro-inflammatory cytokine accumulation | Inflammatory virus | [21] |
| 11 | Allicin | 65036 | Allium sativum (garlic) | Inhibit virus penetration and proliferation (inhibit cell proliferation, protect the heart injury, liver damage, anti-inflammation) | Influenza | [31] |
Table┬Ā3.
Seven antiviral drugs and medicinal value
| No. | Synthetic/Natural drug compound | DB ID | Status | Source/Plant name | Treatment/Property | Reference |
|---|---|---|---|---|---|---|
| 1 | Glycyrrhizic acid (glycyrrhizin) | DB13751 | Approved, experimental | Glycyrrhiza glabra | Inhibit viral replication of SARS-CoV | [23] |
| 2 | Ribavirin | DB00811 | Approved | Synthetic | Effective against chronic hepatitis C virus, SARS-CoV | PMID:18565019, [17] |
| 3 | Tenofovir | DB14126 | Experimental, investigational | Phyllanthus niruri | Hepatitis B virus | [18] |
| 4 | Berberine | DB04115 | Approved, investigational | Berberis vulgaris | Prevents the HIV-PI induced inflammation | [22] |
| 5 | Emodin | DB07715 | Investigational | Radix et Rhizoma Rhei, Radix Polygoni Multiflori | Blocks the S protein of SARS-CoV and ACE2 interaction | [24] |
| 6 | Chloroquine | DB00608 | Approved, investigational, vet approved | Synthetic | HIV, influenza A/H5N1, SARS-CoV, human coronavirus 229E | PMID: 23648708 |
| 7 | Hydroxy chloroquine | DB01611 | Approved | Synthetic | HIV, DENV | PMID:25321315 |
Table┬Ā4.
Docking scores of 18 ligands against SARS-CoV-2 N protein
Table┬Ā5.
Polar interaction (distance Ōēż 3.5 ├ģ) between selected antiviral compounds and nucleocapsid protein of SARS-CoV-2
References
1. Adhikari SP, Meng S, Wu YJ, Mao YP, Ye RX, Wang QZ, et al. Epidemiology, causes, clinical manifestation and diagnosis, prevention and control of coronavirus disease (COVID-19) during the early outbreak period: a scoping review. Infect Dis Poverty 2020;9:29.




2. Jiang S, Hillyer C, Du L. Neutralizing antibodies against SARS-CoV-2 and other human coronaviruses. Trends Immunol 2020;41:355ŌĆō359.



3. Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 2020;11:1620.




4. Wang Q, Zhang Y, Wu L, Niu S, Song C, Zhang Z, et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020;181:894ŌĆō904.



5. McBride R, van Zyl M, Fielding BC. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014;6:2991ŌĆō3018.



7. Lin SM, Lin SC, Hsu JN, Chang CK, Chien CM, Wang YS, et al. Structure-based stabilization of non-native protein-protein interactions of coronavirus nucleocapsid proteins in antiviral drug design. J Med Chem 2020;63:3131ŌĆō3141.



8. Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870ŌĆō1874.




9. Marchler-Bauer A, Anderson JB, Derbyshire MK, DeWeese-Scott C, Gonzales NR, Gwadz M, et al. CDD: a conserved domain database for interactive domain family analysis. Nucleic Acids Res 2007;35:D237ŌĆōD240.



10. Buchan DW, Jones DT. The PSIPRED Protein Analysis Workbench: 20 years on. Nucleic Acids Res 2019;47:W402ŌĆōW407.




11. Huang B. MetaPocket: a meta approach to improve protein ligand binding site prediction. OMICS 2009;13:325ŌĆō330.


12. Rizvi SM, Shakil S, Haneef M. A simple click by click protocol to perform docking: AutoDock 4.2 made easy for non-bioinformaticians. EXCLI J 2013;12:831ŌĆō857.


13. Fuhrmann J, Rurainski A, Lenhof HP, Neumann D. A new Lamarckian genetic algorithm for flexible ligand-receptor docking. J Comput Chem 2010;31:1911ŌĆō1918.


14. Bogusz M, Whelan S. Phylogenetic tree estimation with and without alignment: new distance methods and benchmarking. Syst Biol 2017;66:218ŌĆō231.



15. Sahoo M, Jena L, Rath SN, Kumar S. Identification of suitable natural inhibitor against influenza A (H1N1) neuraminidase protein by molecular docking. Genomics Inform 2016;14:96ŌĆō103.



16. Moghadamtousi SZ, Kadir HA, Hassandarvish P, Tajik H, Abubakar S, Zandi K. A review on antibacterial, antiviral, and antifungal activity of curcumin. Biomed Res Int 2014;2014:186864.


17. Ashfaq UA, Idrees S. Medicinal plants against hepatitis C virus. World J Gastroenterol 2014;20:2941ŌĆō2947.
18. Mohan M, James P, Valsalan R, Nazeem PA. Molecular docking studies of phytochemicals from Phyllanthus niruri against hepatitis B DNA polymerase. Bioinformation 2015;11:426ŌĆō431.



19. Doki T, Tarusawa T, Hohdatsu T, Takano T. In vivo antiviral effects of U18666A against type I feline infectious peritonitis virus. Pathogens 2020;9:67.

20. Alhazmi MI. Molecular docking of selected phytocompounds with H1N1 proteins. Bioinformation 2015;11:196ŌĆō202.



21. Rafe T, Shawon PA, Salem L, Chowdhury NI, Kabir F, Bin Zahur SM, et al. Preventive role of resveratrol against inflammatory cytokines and related diseases. Curr Pharm Des 2019;25:1345ŌĆō1371.



22. Kaliyaperumal S, Periyasamy K, Balakrishnan U, Palanivel P, Egbuna C. Antiviral phytocompounds for drug development: a data mining studies. In: Phytochemicals as Lead Compounds for New Drug Discovery (Egbuna C, Kumar S, Ifemeje J, Ezzat S, Kaliyaperumal S, eds.). Amsterdam: Elsevier, 2020. pp. 239-244.


23. Chen F, Chan KH, Jiang Y, Kao RY, Lu HT, Fan KW, et al. In vitro susceptibility of 10 clinical isolates of SARS coronavirus to selected antiviral compounds. J Clin Virol 2004;31:69ŌĆō75.



24. Ho TY, Wu SL, Chen JC, Li CC, Hsiang CY. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antiviral Res 2007;74:92ŌĆō101.



25. Jagadeb M, Rath SN, Sonawane A. In silico discovery of potential drug molecules to improve the treatment of isoniazid-resistant Mycobacterium tuberculosis. J Biomol Struct Dyn 2019;37:3388ŌĆō3398.



26. Kadioglu O, Saeed M, Johannes Greten H, Efferth T. Identification of novel compounds against three targets of SARS CoV-2 coronavirus by combined virtual screening and supervised machine learning. Preprint at http://dx.doi.org/10.2471/BLT.20.255943 (2020).


27. Calligari P, Bobone S, Ricci G, Bocedi A. Molecular investigation of SARS-CoV-2 proteins and their interactions with antiviral drugs. Viruses 2020;12:445.



28. Sarma P, Shekhar N, Prajapat M, Avti P, Kaur H, Kumar S, et al. In-silico homology assisted identification of inhibitor of RNA binding against 2019-nCoV N-protein (N terminal domain). J Biomol Struct Dyn 2020 May 18 [Epub]. https://doi.org/10.1080/07391102.2020.1753580.



29. Bhowmik D, Nandi R, Jagadeesan R, Kumar N, Prakash A, Kumar D. Identification of potential inhibitors against SARS-CoV-2 by targeting proteins responsible for envelope formation and virion assembly using docking based virtual screening, and pharmacokinetics approaches. Infect Genet Evol 2020;84:104451.



30. Kang S, Yang M, Hong Z, Zhang L, Huang Z, Chen X, et al. Crystal structure of SARS-CoV-2 nucleocapsid protein RNA binding domain reveals potential unique drug targeting sites. Acta Pharm Sin B 2020;10:1228ŌĆō1238.


31. Johns C. Glycyrrhizic acid toxicity caused by consumption of licorice candy cigars. CJEM 2009;11:94ŌĆō96.
- TOOLS
-
METRICS

-
- 10 Crossref
- 0 Scopus
- 6,738 View
- 113 Download
- Related articles in GNI







